Sequencing of first frog genome sheds light on treating disease
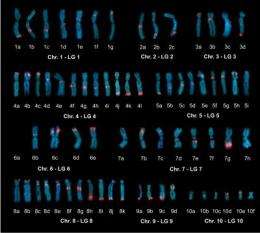
A pair of University of Houston researchers contributed to the assembly of the first comprehensive DNA sequence of an amphibian genome, which will shed light on the study of embryonic development, with implications for preventing birth defects and more effectively treating many human diseases.
Amy Sater and Dan Wells, both professors in UH's department of biology and biochemistry, collaborated with a number of other scientists in what Sater calls "a massive and international effort," landing them a cover story - "The Genome of the Western Clawed Frog Xenopus tropicalis" - in a recent issue of Science magazine, the world's leading journal of original scientific research, global news and commentary.
Originating in West Africa, Xenopus tropicalis is a frog that is extremely important for studies of embryonic development and the regulation of cell division. The genes in frogs are highly similar to those in mice and humans, as are the key communication pathways. These molecular communication pathways serve as lines of communication between cells and are critical to control how cells choose to form the brain, limbs, muscle cells and the pancreas. They also are important for the maintenance and differentiation of stem cells, including those that maintain the lining of our intestines. Many experiments can be carried out in Xenopus more quickly and easily, as well as far less expensively, than in mouse embryos, and the tools provided by the genome assembly will transform research using this animal.
"In many cases, if one of these key communication pathways is misregulated due to a key gene being mutated, it can lead to several major types of cancer," Sater said. "This particular frog is a terrific animal in which to study these pathways because you can study both the biochemistry of how the pathways work, as well as what the pathway is actually doing in developing embryos.
"Working out the biochemical mechanism is extremely difficult to do in a mouse embryo. We can obtain hundreds of these frog embryos that are developing synchronously, and because they are fertilized and develop outside the mother, we can watch and manipulate specific events much more easily and on a much larger scale than in mouse embryos."
Sater and Wells' contributions were in the difficult process of assembly, after they collaborated with scientists from the Human Genome Sequencing Center at the Baylor College of Medicine to generate a genetic map. The project is funded by a nearly $2 million grant from the National Institutes of Health. Ultimately, Sater likened sequencing a genome to assembling a 10,000-piece jigsaw puzzle without having a detailed picture from which to work. The genetic map prepared by Sater and her colleagues provided a big part of that picture to guide long-range assembly of the puzzle.
Once the UH and Baylor team's portion was complete, they compared the short sequences used as landmarks in their genetic map with the genome sequences. These comparisons allowed their colleagues at University of California, Berkeley, to complete the assembly of the genome.
"Sequencing and assembling a genome is basically science infrastructure - the equivalent of building roads and bridges - and once the infrastructure is in place, everyone can benefit," Sater said. "This work is an enormous contribution to research now in progress throughout the world, and essentially every study that uses Xenopus as a research animal gets a big boost from this project."
Big science like this, Sater said, has a lot of authors and provides fundamental, important information for all biologists in trying to understand how specific genes function. Important contributions also came from individuals at the Joint Genome Institute, Cambridge, University of California Irvine, Washington University School of Medicine, University of Virginia, the National Institutes of Health, the Université d'Evry in France, the National Institute for Medical Research in the United Kingdom and the Okinawa Institute for Science and Technology in Japan.
"Many human diseases, such as cancer, heart disease and hereditary conditions, can be traced back to changes in how genes are expressed, and it may be possible to treat these and other diseases more effectively if we understand how these genes function and how they are turned on and off," Sater said. "Having this blueprint provides us with landmarks that we can use to change when and where certain genes are expressed. The toolkit provided by this study will allow us to examine the functions of individual genes that have already been identified as key players in specific events and important to human health."
Provided by University of Houston
















