8.2 percent of our DNA is 'functional'
Only 8.2% of human DNA is likely to be doing something important – is 'functional' – say Oxford University researchers.
This figure is very different from one given in 2012, when some scientists involved in the ENCODE (Encyclopedia of DNA Elements) project stated that 80% of our genome has some biochemical function.
That claim has been controversial, with many in the field arguing that the biochemical definition of 'function' was too broad – that just because an activity on DNA occurs, it does not necessarily have a consequence; for functionality you need to demonstrate that an activity matters.
To reach their figure, the Oxford University group took advantage of the ability of evolution to discern which activities matter and which do not. They identified how much of our genome has avoided accumulating changes over 100 million years of mammalian evolution – a clear indication that this DNA matters, it has some important function that needs to be retained.
'This is in large part a matter of different definitions of what is "functional" DNA,' says joint senior author Professor Chris Pointing of the MRC Functional Genomics Unit at Oxford University. 'We don't think our figure is actually too different from what you would get looking at ENCODE's bank of data using the same definition for functional DNA.
'But this isn't just an academic argument about the nebulous word "function". These definitions matter. When sequencing the genomes of patients, if our DNA was largely functional, we'd need to pay attention to every mutation. In contrast, with only 8% being functional, we have to work out the 8% of the mutations detected that might be important. From a medical point of view, this is essential to interpreting the role of human genetic variation in disease.'
The researchers Chris Rands, Stephen Meader, Chris Ponting and Gerton Lunter report their findings in the journal PLOS Genetics. They were funded by the UK Medical Research Council and the Wellcome Trust.
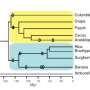
The researchers used a computational approach to compare the complete DNA sequences of various mammals, from mice, guinea pigs and rabbits to dogs, horses and humans.
Dr Gerton Lunter from the Wellcome Trust Centre for Human Genetics at Oxford University, the other joint senior author, explained: 'Throughout the evolution of these species from their common ancestors, mutations arise in the DNA and natural selection counteracts these changes to keep useful DNA sequences intact.'
The scientists' idea was to look at where insertions and deletions of chunks of DNA appeared in the mammals' genomes. These could be expected to fall approximately randomly in the sequence – except where natural selection was acting to preserve functional DNA, where insertions and deletions would then lie further apart.
'We found that 8.2% of our human genome is functional,' says Dr Lunter. 'We cannot tell where every bit of the 8.2% of functional DNA is in our genomes, but our approach is largely free from assumptions or hypotheses. For example, it is not dependent on what we know about the genome or what particular experiments are used to identify biological function.'
The rest of our genome is leftover evolutionary material, parts of the genome that have undergone losses or gains in the DNA code – often called 'junk' DNA.
'We tend to have the expectation that all of our DNA must be doing something. In reality, only a small part of it is,' says Dr Chris Rands, first author of the study and a former DPhil student in the MRC Functional Genomics Unit at Oxford University.
Not all of the 8.2% is equally important, the researchers explain.
A little over 1% of human DNA accounts for the proteins that carry out almost all of the critical biological processes in the body.
The other 7% is thought to be involved in the switching on and off of genes that encode proteins – at different times, in response to various factors, and in different parts of the body. These are the control and regulation elements, and there are various different types.
'The proteins produced are virtually the same in every cell in our body from when we are born to when we die,' says Dr Rands. 'Which of them are switched on, where in the body and at what point in time, needs to be controlled – and it is the 7% that is doing this job.'
In comparing the genomes of different species, the researchers found that while the protein-coding genes are very well conserved across all mammals, there is a higher turnover of DNA sequence in the regulatory regions as this sequence is lost and gained over time.
Mammals that are more closely related have a greater proportion of their functional DNA in common.
But only 2.2% of human DNA is functional and shared with mice, for example – because of the high turnover in the regulatory DNA regions over the 80 million years of evolutionary separation between the two species.
'Regulatory DNA evolves much more dynamically that we thought,' says Dr Lunter, 'but even so, most of the changes in the genome involve junk DNA and are irrelevant.'
He explains that although there is a lot of functional DNA that isn't shared between mice and humans, we can't yet tell what is novel and explains our differences as species, and which is just a different gene-switching system that achieves the same result.
Professor Ponting agrees: 'There appears to be a lot of redundancy in how our biological processes are controlled and kept in check. It's like having lots of different switches in a room to turn the lights on. Perhaps you could do without some switches on one wall or another, but it's still the same electrical circuit.'
He adds: 'The fact that we only have 2.2% of DNA in common with mice does not show that we are so different. We are not so special. Our fundamental biology is very similar. Every mammal has approximately the same amount of functional DNA, and approximately the same distribution of functional DNA that is highly important and less important. Biologically, humans are pretty ordinary in the scheme of things, I'm afraid.
'I'm definitely not of the opinion that mice are bad model organisms for animal research. This study really doesn't address that issue,' he notes.
More information: PLoS ONE DOI: 10.1371/journal.pgen.1004525
Journal information: PLoS Genetics , PLoS ONE
Provided by Oxford University