Scientists develop potential new drug treatment to tackle viruses
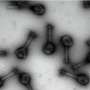
(Phys.org) —An international team of scientists have successfully developed a novel compound which early signs suggest might prevent a range of viruses from infecting humans. Researchers from Oxford, Beijing, Leeds and Innsbruck collaborated on the new inhibitor. It targets a group of viruses responsible for hand, foot and mouth disease, especially the EV71 virus. This viral group causes numerous epidemics in children, mainly in Asia, with roughly 10 million cases reported every year in China alone. Symptoms are usually mild but in some cases the disease can prove fatal - the Chinese government reported over 900 deaths in 2010. The disease is currently untreatable and is a major global threat to public health.
This discovery, published in Nature Structural and Molecular Biology, may also have important implications for combating other diseases. Hand, foot and mouth disease is caused by several closely related viruses, and the new compound is effective against all of these. In addition, other wide-spread viruses are quite closely related (they belong to the same enterovirus genus), including poliovirus and many of the viruses responsible for the common cold.
Professor Dave Stuart, Director of the Division of Structural Biology (STRUBI) in the Nuffield Department of Medicine at the University of Oxford and Life Science Director at Diamond, who worked on the new inhibitor, explains: "By targeting a structural feature also found in related viruses, it should be possible to devise similar therapeutics to target them. Within the field, I am aware of one company that is already making progress in targeting the major common cold virus. Our work is still at an early stage, but we are working with academic groups in China to take the inhibitor forward."
The new drug treatment, which so far has been tested only in isolated cells, was created using cutting edge techniques. Structure-based drug design refers to drug treatments that are specifically designed to target the structure of a disease; it works in three stages. Firstly the pathogen's structure must be solved. Secondly, scientists must determine how that structure relates to the functioning of the pathogen. Finally, they can develop a means of interacting with that structure to disable the pathogen.
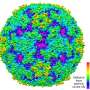
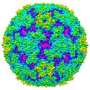
The structure of EV71 was first solved at Diamond Light Source, the UK's national synchrotron, less than 2 years ago, in 2012. After this, scientists used the giant microscope to determine how the virus changes shape to enter the host. Enteroviruses are part of a larger family of viruses called picornaviruses. These viruses have an outer shell which, once inside a host cell, breaks apart releasing the viral RNA into the cell. The team have been able to discover a very potent molecule which disables the mechanism by which the viral shell breaks open, thus stopping the virus in its tracks by making it unable to infect the host.
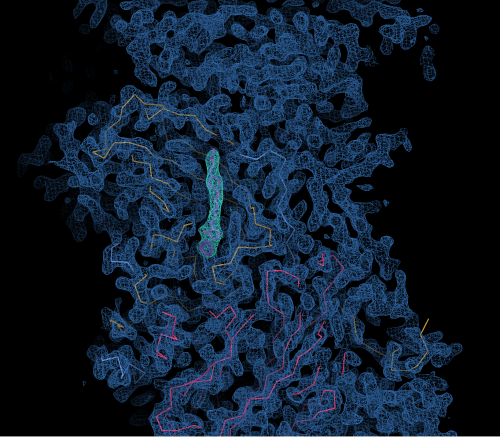
The Anglo-Chinese collaboration used two of Diamond's life science beamlines, I03 and I24, to develop the new methodology. When they determined the virus structure in 2012, the team saw a small pocket inside the virus where a drug might fit. They have now developed a potential drug molecule that can sit inside this pocket, where it locks the virus rigid, preventing it breaking open and releasing its RNA.
This research is still in its early stages, and there is a considerable way to go before the drug treatment is available on the market. It's also important to note that viruses evolve exceptionally fast, and it may be that EV71 mutates to overcome the inhibitor. However, the early signs are promising, and now that the methodology exists, scientists may be able to develop the inhibitor to become resistant to mutations. The speed with which this development has come about – in less than two years from the original discovery of the virus structure – demonstrates the vast potential of structure-based drug design, and the remarkable capabilities that are now at scientists' disposal in the fight against disease.
More information: 'More-powerful virus inhibitors from structure-based analysis of HEV71 capsid-binding molecules' will be published in Nature Structural & Molecular Biology. DOI: 10.1038/nsmb.2769
Journal information: Nature Structural & Molecular Biology , Nature Structural and Molecular Biology