Tracing the protein assembly line
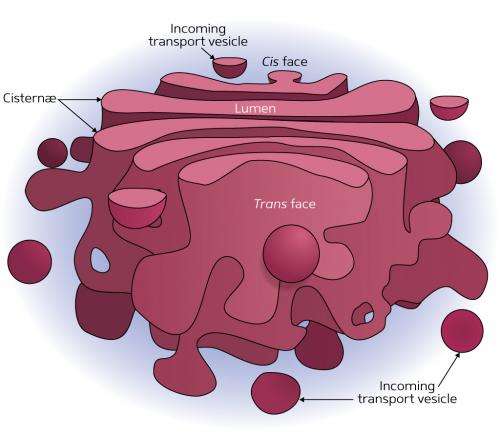
Many newly synthesized proteins undergo a sequence of enzymatic modifications that enable them to do their jobs better. This process occurs within a series of membrane-bound structures called 'cisternae' that form the Golgi apparatus—an organelle that packages proteins before they are sent to their destination. Akihiko Nakano, Yasuyuki Suda and colleagues at the RIKEN Center for Advanced Photonics have now gained important insights into how protein movement through the Golgi apparatus is regulated.
When new proteins are produced, they are deposited into cisternae on the 'cis' side of the Golgi. These proteins undergo a maturation process as they migrate toward the 'trans' side of the Golgi before being released into the cell as fully processed proteins (Fig. 1). Nakano's team previously characterized this Golgi maturation process in yeast cells, resolving a long-standing debate in cell biology. "However, the molecular machinery of this cisternal maturation remained a mystery," says Suda.
Nakano, Suda and their colleagues are interested in enzymes known as Rab GTPases, which help to coordinate molecular trafficking within the cell. Different Rab GTPase proteins can serve as 'bar codes' that specifically identify cisternae in different zones of the Golgi. For example, the protein Ypt1 is found in the cis-Golgi in yeast cells, whereas Ypt32 resides in the trans-Golgi. "It has been speculated that the transition of Rab GTPases drives Golgi maturation because the transition from Ypt1 to Ypt32 coincides with this process," says Suda.
The researchers set out to determine the role of a third Rab GTPase, Ypt6. They found that Ypt6 generally resides in the middle region of the Golgi, and showed that although Ypt6 can be found alongside Ypt1, it is generally absent from Ypt32-bearing cisternae in the trans-Golgi. Rab GTPases are only bound to membranes when they are switched on. Subsequent experiments revealed that Ypt32 actively recruits another enzyme that turns Ypt6 off, thereby facilitating its removal from the membrane as cisternal maturation proceeds.
However, the absence of this Rab transition does not impair cell survival, indicating that this protein is not quite the last piece of this puzzle. "The resulting defects only made maturation a bit slower than usual, suggesting that other or perhaps multiple regulatory mechanisms drive Golgi maturation," says Suda. To delve more deeply into these processes, Nakano's lab has devised an innovative microscopy platform that allows the researchers to simultaneously observe multiple proteins in living cells in detail. "Using this device, we'd like to clarify the fundamental molecular machinery underlying protein trafficking."
More information: Suda, Y., Kurokawa, K., Hirata, R. & Nakano, A. Rab GAP cascade regulates dynamics of Ypt6 in the Golgi traffic. Proceedings of the National Academy of Sciences USA 110, 18976–18981 (2013). dx.doi.org/10.1073/pnas.1308627110
Journal information: Proceedings of the National Academy of Sciences
Provided by RIKEN