Comprehensive new map details dynamics of gene activity during embryonic development

Embryonic development involves the tightly coordinated activity of thousands of genes, each switched on at a specific time and place in the growing organism under the control of regulatory DNA sequences called promoters. An international team of researchers including Piero Carninci from the RIKEN Center for Life Science Technologies has now produced the first vertebrate 'promoterome', or genome-wide map of how promoter usage changes during development.
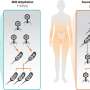
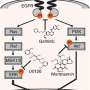
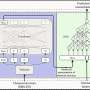
Carninci and his colleagues obtained RNA samples from zebrafish at 12 key stages of development and used a technique called cap analysis of gene expression (CAGE) to determine promoter usage in each sample. The samples contain messenger RNA transcripts—temporary copies of DNA sequences that carry the information for protein synthesis—and other long non-coding RNAs. Transcripts can be identified and isolated by virtue of a common motif called the 5' cap and then sequenced to establish which promoter was used to synthesize them.
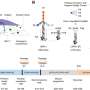
Analysis of the data revealed a number of hitherto unknown features of gene regulation. After the very early stages of development, RNAs inherited from the mother are destroyed and substituted by RNAs produced by the embryo. These RNAs are produced by a different set of promoters than those involved in the production of maternal transcripts. This is the first time that such a switch in promoter usage has been mapped at a broad scale and with such high accuracy.
The researchers compared the data to equivalent datasets collected previously for fruit flies, pufferfish and humans. The comparison suggests that all vertebrates have remarkably similar gene regulation mechanisms, which are fundamentally different from those of invertebrates.
They also identified a previously unknown regulatory sequence within vertebrate promoters. This sequence, which is rich in the DNA bases adenine and guanine, is likely to be a common regulatory element in all vertebrate species.
The zerbrafish promoterome is the first large-scale map of any fish genome, and the results suggest that there are important features that are conserved in the basic structure of the promoters. "We identified short stretches of DNA that are used in the same way in distant genomes," says Carninci. "In particular, the elements that cause the gene to initiate RNA transcription."
Carninci believes that the application of CAGE will be highly beneficial for the identification of regulatory DNA sequences in many organisms. "We are now expanding the utilization of CAGE to several other organisms," he notes.
More information: Nepal, C., et al. Dynamic regulation of the transcription initiation landscape at single nucleotide resolution during vertebrate embryogenesis, Genome Research advance online publication, 3 September 2013 (DOI: 10.1101/gr.153692.112). dx.doi.org/10.1101/gr.153692.112
Journal information: Genome Research
Provided by RIKEN