Debate ends: Team reconciles puzzling findings relating to centromere structure
Scientists at the Stowers Institute of Medical Research have developed an innovative method to count the number of fluorescent molecules in a cluster and then applied the novel approach to settle a debate rampant among cell biologists—namely, how DNA twists into a unique chromosomal structure called the centromere. Knowing this helps explain how cells navigate the hazards of division and avoid the disastrous consequences of ending up with the wrong number of chromosomes.
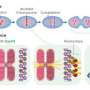
Centromeres, which sit at the cross point of the "X" used to represent duplicated chromosomes, are DNA structures that link those duplicated strands when cells are poised to divide. As division starts, a complex cellular machine drags each chromosome to opposite poles of the cell by grabbing onto centromeres and pulling each arm of the "X" into what will become a daughter cell.
Researchers had known that a nucleosome—a short coil of DNA twisted around a core of proteins—forms at each centromere. Within the core is a protein, called Cse4 in yeast, that is found only at that location. But the overall architecture of that nucleosome was unknown. Now, Stowers Associate Investigator Jennifer Gerton, Ph.D., has used live cell imaging to reveal constituents of the centromeric protein core. That study is published in the July 20, 2012 of the journal Cell.
"Understanding centromeres is critical because of the role they play in maintaining genomic integrity," says Gerton. "Losing a chromosome is catastrophic for any cell. And if it happens in sperm or egg cells, it is associated with conditions like Down's Syndrome."
Gerton, whose lab uses both the yeast Saccharomyces cerevisiae and mammalian cells to study the mechanics of cell division, says that, previously, people had proposed at least 6 different centromere structures. "What we found is that centromeric nucleosomes change their structure during cell division," she says. "That explained why people had observed different structures. They had likely been looking at different phases of the cell cycle."
"By demonstrating a new method for monitoring the composition of centromeric nucleosomes in living cells, this work helps to resolve some of the controversies surrounding the architecture of the centromere," said Anthony Carter, Ph.D., of the National Institutes of Health's National Institute of General Medical Sciences, which partially funded the research. "The findings have important implications for understanding chromosome segregation, and may lead to insights on how the process goes awry in certain genetic diseases."
Aiding the effort were Stowers Research Advisors Jay Unruh, Ph.D., and Brian Slaughter, Ph.D., who combined two microscopy methods to probe yeast cells engineered to express Cse4 hooked to a green fluorescent protein (GFP) tag. The approach allowed them to track and then count in a living cell the number of Cse4 molecules in a centromeric nucleosome, a question hotly debated in the field.
Slaughter describes the controversy more prosaically: "To a microscopist, the question came down to, how many GFP molecules can we see in a fluorescent dot in the middle of a yeast cell?"
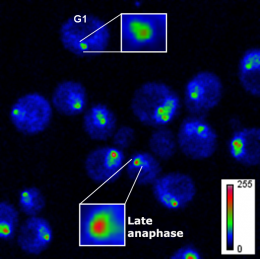
Although the microscopy technology applied—fluorescence correlation spectroscopy coupled with calibrated imaging—sounds and is complicated, doing the math required to settle the controversy hardly required a calculator. Yeast cells have 16 chromosomes, each with one centromere. If each centromere contained just one copy of Cse4, then the dot glowing in each cell should be 16 times brighter than a single GFP molecule. And it was. But only right before cells began dividing. Once chromosomes separated and moved to opposite poles of a dividing cell, a stage biologists call anaphase, the intensity of the signal increased.
"To our surprise, we quickly realized that we observed 16 Cse4-GFP molecules early in the cell cycle, and then 32 Cse4-GFP molecules in anaphase," says Slaughter. "That meant the composition of the complex was changing." Further analysis indicated that as cells moved into anaphase a component of the centromeric nucleosome got booted out of the core complex and was replaced by an extra molecule of Cse4, changing both the shape and size of the centromere.
Gerton's team, led by the study's first author Manjunatha Shivaraju, Ph.D., confirmed these findings using additional approaches. They found evidence that two molecules of Cse4 were interacting at the centromere in anaphase, but these interactions were not present during the rest of the cell cycle.
This work will be published back to back with a parallel study of human cells by Yamimi Dalal, Ph.D., of the National Cancer Institute. "The timing of structural changes differs in yeast and human cells," Gerton says, referring to the human study. "And we visualized nucleosomes differently than the Dalal group did. But our conclusions are the same—that human and yeast centromeres undergo similar dynamic changes with the cell cycle."
Unruh and Slaughter, who developed the microscopy approach used in the study, act as in-house consultants to Stowers investigators about molecular imaging. "Research Advisors provide collaborative support for projects requiring particular expertise," says Slaughter. "We develop novel ways to address questions the PIs are asking." (Stowers also employs research advisors specializing in mathematical modeling and genomics.)
Why such massive effort should be expended on centromere components is evident, given the disastrous consequences of cell division errors. "Most cancer cells are aneuploid," says Shivaraju, referring to a condition in which cells exhibit abnormal numbers of chromosomes. "Knowing that centromeres undergo this structural oscillation could tell us how aneuploidy occurs at a molecular level."
Gerton concurs but also sees the work as reinforcing the utility of yeast as a model organism. "The fact that nucleosome structure is conserved between humans and yeast shows that yeast is a fantastic model for studying molecular mechanisms underlying cell division," she says. "We will continue to use yeast to understand factors that trigger structural changes we see in centromeric nucleosomes."
Journal information: Cell
Provided by Stowers Institute for Medical Research