A new way of looking at photosystem II: Researchers study key protein complex crucial to photosynthesis
Future prospects for clean, green, renewable energy may hinge upon our ability to mimic and improve upon photosynthesis – the process by which green plants, algae and some bacteria convert solar energy into electrochemical energy. An artificial version of photosynthesis, for example, could use sunlight to produce liquid fuels from nothing more than carbon dioxide and water. First, however, scientists need a better understanding of how a large complex of proteins, called photosystem II, is able to split water molecules into oxygen, electrons and hydrogen ions (protons). A new road to reaching this understanding has now been opened by an international team of researchers, led by scientists at the Lawrence Berkeley National Laboratory (Berkeley Lab) and SLAC National Accelerator Laboratory.
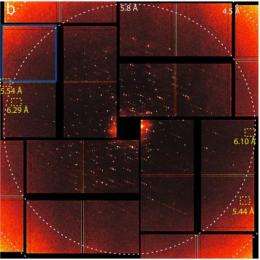
Using ultrafast, intensely bright pulses of X-rays from SLAC's Linac Coherent Light Source (LCLS), the research team produced the first ever images at room temperature of microcrystals of the photosystem II complex. Previous imaging studies, using X-rays generated via synchrotron radiation sources, required cryogenic freezing, which alters the samples. Also, to catalyze its reactions, photosystem II relies upon an enzyme that contains a manganese-calcium cluster that is highly sensitive to radiation. With the high-intensity femtosecond X-ray pulses of the LCLS, the research team was able to record intact images of these clusters before the radiation destroyed them.
"We have demonstrated that the 'probe before destroy' strategy of the LCLS is successful even for the highly-sensitive oxygen bridged manganese-calcium cluster in photosystem II at room temperature," says Vittal Yachandra, a chemist with Berkeley Lab's Physical Biosciences Division. "This is an important step toward future studies for resolving the composition and atomic structure of the manganese-calcium cluster in the photosystem II complex during the critical formation of oxygen molecules."
Yachandra is one of the corresponding authors of a paper describing this work in the Proceedings of the National Academy of Sciences (PNAS) titled "Room temperature femtosecond X-ray diffraction of photosystem II microcrystals. The other corresponding authors are Junko Yano, also a chemist with Berkeley Lab's Physical Biosciences Division, and Uwe Bergmann of SLAC.
For more than two billion years, nature has employed photosynthesis to oxidize water into molecular oxygen. Photosystem II, the only known biological system that can harness visible light for the photooxidation of water, produces most of the oxygen in Earth's atmosphere through a five-step catalytic cycle (S0-to-S4 oxidation states). Light-harvesting proteins in the complex capture solar photons that energize the manganese-calcium cluster and drive a series of oxidations and proton transfers that in the final S4 state forms the bond between oxygen atoms that yields molecular oxygen.
Scientific teams in the past, including several led by Yachandra and Yano, have shed much light on the S0 through S3 oxidation states of the manganese-calcium cluster, which remain stable for several seconds. However, the S4 state is highly reactive and has not yet been fully characterized in experiments.
"Capturing the S4 state in a time-resolved manner will be essential for understanding the water-oxidation mechanism," Yano says. "While X-ray diffraction is clearly the technique of choice for such detailed structural studies, the inherent radiation sensitivity of the manganese-calcium cluster poses a major challenge for protein crystallography on synchrotron radiation sources."
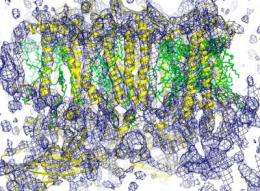
SLAC's LCLS is an X-ray laser powered by a two-mile-long linear accelerator (or linac) that generates pulses of X-ray light on a femtosecond timescale. These pulses are more than a billion times brighter than those from the most powerful synchrotrons. Yachandra, Yano and their colleagues suspended photosystem II microcrystals in a liquid that was jet-streamed into the path of the pulsed light. The diffraction of LCLS X-rays passing through the photosystem II microcrystals created patterns that computers could reconstruct into images of the complex's composition and atomic structure at a resolution of 6.5 angstroms - one ten-billionth of a meter or about the diameter of a hydrogen atom.
"We hope that with improved samples, in the future we will be able to get to a higher resolution – perhaps 3 angstroms or better," says Jan Kern, a research scientist at Berkeley Lab and SLAC who was the lead author on the PNAS paper.
Photosystem II microcrystals (approximately 10 micrometers in diameter)were used as a matter of efficiency. Molecular reconstruction through X-ray diffraction requires the examination of literally millions of crystals, since each shot from the LCLS destroys the specimen.
"Because it takes months to grow sufficient quantities of the photosystem II complex in bacterial culture, the use of microcrystals made the most efficient use of time and materials," says Kern. "Also, microcrystals were much easier to direct toward the LCLS X-ray beam using the liquid-stream sample delivery system developed by our collaborators at SLAC."
Paul Adams and Nicholas Sauter, also with Berkeley Lab's Physical Biosciences Division and also co-authors of the PNAS paper, led the data analysis in this study, writing new software to manage the computations.
"Doing this study was a monumental achievement that required a large team to make it happen," Sauter says. "We injected crystal samples into the beam at a rate of 120 per second, and after a week we had 63 Terabytes of data from which we selected the best 7,000 diffraction images to reconstruct photosystem II's molecular structure."
Further studies at the LCLS by the research team are already underway using both X-ray diffraction and spectroscopy techniques to investigate the intermediate reaction states formed in the photosystem II complex as it undergoes photooxidation.
"We hope to learn from nature's design principles and apply that knowledge to the design and development of artificial photosynthetic systems," Yano says.
Journal information: Proceedings of the National Academy of Sciences
Provided by Lawrence Berkeley National Laboratory




















