A cell's first steps: Building a model to explain how cells grow
A collaboration between Lehigh University physicists and University of Miami biologists addresses an important fundamental question in basic cell biology: How do living cells figure out when and where to grow?
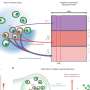
The teams of Assistant Professor Dimitrios Vavylonis and Associate Professor Fulvia Verde discovered that protein Cdc42 oscillates throughout yeast cells, precipitating a ballet of proteins that change its polarity. By changing polarity, Cdc42 regulates shape, structure and function in yeast cells, starting the growth process by clustering in an area of the cellular membrane. The oscillatory mechanism they found may be a general strategy among all self-organizing biological systems, not just simple yeast.
"The research is fundamental because it provides science with an important answer to how a living cell controls its growth process," said Vavylonis. "Knowing how this particular protein controls growth could in the long run affect the search for drugs to control cell growth for tissue regeneration, organ development, and explain how neurons extend in different directions."
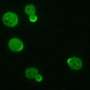
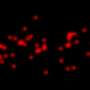
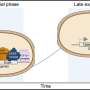
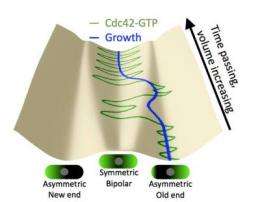
This work indicates how Cdc42 activates bipolar growth only once a minimal cell length has been achieved. At that point, Cdc42 begins to oscillate back and forth through the cell, as the two tips compete for it. Using fluorescent markers to tag each of the many proteins involved, researchers observed the Cdc42 protein oscillate from side to side within a cell, switching sides about every five minutes. The fluctuations provide an adaptable mechanism for cells to control their size and structure in the fast-changing environment within.
The study, Oscillatory Dynamics of Cdc42 GTPase In The Control of Polarized Growth, appears today in the journal Science.
The findings demonstrate just part of the complex process of cell growth and differentiation, but mark how advanced the science of biophysics has become. Only recently has the clear imaging and monitoring of protein activity become possible at the minute sizes and shortened time scales of individual cell maturation.

"Up until now, no one has ever seen the way this protein oscillates back and forth throughout the cell," said Tyler Drake, a Lehigh graduate student and co-author of the paper. "Looking at a simple system like yeast may allow us to understand the principles behind growth in other cells."
The Lehigh team developed the mathematical model of this phenomenon by analyzing cell data collected by Maitreyi Das and Fulvia Verde at the University of Miami. Drake and Vavylonis used a Lehigh Class of 1968 Junior Faculty Fellowship and a Sigma Xi grant to visit the University, where they began to test their mathematical theory. According to the model, changes in abundance or activity of Cdc42, or of its regulators, can shift the system to more asymmetric or symmetric states. The model's conclusions were supported by biological observations of the Miami team, who genetically manipulated regulators of the protein and realized they could change cell shape and growth symmetry by adjusting Cdc42.
Vavylonis's research has for years explored the way the cellular cytoskeleton organizes and functions. In collaboration with biologists and computer scientists, his team uses physics to study, analyze, and model the physical properties of these adaptive biological materials.
Journal information: Science
Provided by Lehigh University