Scientists devise new imaging technique for analysis of biological samples
(PhysOrg.com) -- When trying to understand how cells respond to toxins, scientists want to do as little sample preparation as possible. Preparing these cells by immersing them in chemicals or drying them out can erase vital information. At Pacific Northwest National Laboratory, scientists proved that a new ionization technique they developed in 2009 can provide fingerprint and locate proteins, amino acids, and other chemicals in cells that make up tissues or microbial communities using mass spectrometry.
"The beauty of the technique is that it doesn't require any sample preparation," said Dr. Julia Laskin, who led the research project for PNNL's Chemical Imaging Initiative. "Here, you just take your sample, slice it, and put it in front of the instrument for analysis."
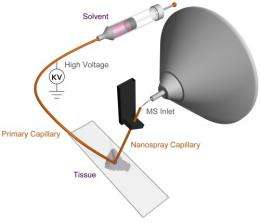
The technique, known as Nanospray Desorption Electrospray Iionization or nano-DESI, allows scientists to efficiently determine which molecules reside in a precise spot on a sample. With this information, researchers can learn more about how diverse biological samples, such as tissues and microbes, respond to environmental factors. For example, biochemists can gain see how the marine microbe Shewanella oneidensis alter metals to remediate hazardous materials in the soil. For example, S. oneidensis can change very soluble hexavalent uranium to less soluble form, limiting its movement in groundwater. Another opportunity lies with medical researchers learning how nicotine and other toxins affect brain cells.
"Great discoveries often require great tools," said Dr. Louis Terminello, who leads the Chemical Imaging Initiative at PNNL. "The discoveries needed to solve today's problems aren't something that you're going to get by eyeballing a sample."
From the beginning, the team was convinced that the liquid bridge used in nano-DESI could be scaled down to analyze small areas on biological samples. By making adjustments, the team was able to scale the probe down to analyze an area about 10 micrometers in diameter, about the same size as a single red blood cell or mid-sized bacteria.
"With this probe, we are getting down to individual cells," said Laskin.
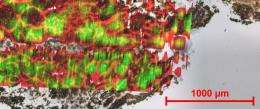
The team first analyzed rat brain tissues, which provide an outstanding test case for the technique, because they are very dense and yield high signals. The nano-DESI was able to draw up and analyze the molecules from different regions on the sample. Then, the team moved onto "airy" kidney tissues, and again were able to analyze micrometer-sized areas.
With each sample, the nanoDESI generated reams of mass spectra. Existing software packages could not process the data. The job fell to Brandi Heath, a team member who recently completed her bachelor's degree and is working on her master's degree at Washington State University. She read the charts and determined the fatty acids, amino acids, lipids, and other molecules that resided at different locations on the tissue.
"It was very tedious, but in the end, very rewarding," said Heath.
"Compared to other online liquid extraction techniques, nano-DESI has about an order of magnitude better spatial resolution," Laskin added. "It is comparable, spatially, to what laser-based techniques can give."
The team is working on two efforts related to their work with nanoDESI. First, they are working with Drs. Dongsheng Li and James Carson through the Chemical Imaging Initiative to understand and visualize the mass spectrometry data on the fly. Also, the team is working with Drs. Matthew Marshall, Margaret Romine, Grigoriy Pinchuk, and Jim Fredrickson to analyze different microbial communities of interest to the Department of Energy.
More information: Laskin J, et al. 2012. "Tissue Imaging Using Nanospray Desorption Electrospray Ionization Mass Spectrometry." Analytical Chemistry 84(1):141-148. DOI: 10.1021/ac2021322
Journal information: Analytical Chemistry
Provided by Pacific Northwest National Laboratory