Scientists study protein dynamical transitions
(PhysOrg.com) -- Central to life and all cellular functions, proteins are complex structures that are anything but static, though often illustrated as two-dimensional snapshots in time.
Cornell scientists, using the Cornell High Energy Synchrotron Source (CHESS), have gained new insight into the underlying mechanisms of how protein structures change at low temperatures. Their paper, whose first author is CHESS staff scientist Chae Un Kim, was published online Dec. 12 in Proceedings of the National Academy of Sciences. Kim's co-authors are Mark Tate, senior research associate in the Laboratory of Atomics and Solid State Physics; and Sol Gruner, professor of physics and CHESS director.
Proteins fluctuate in biological functions ranging from enzymatic catalysis to interactions with other molecules, including DNA. When they are cooled, the fluctuations dampen and eventually stop, typically between -70 and -30 degrees Celsius (-94 to -22 Fahrenheit); this is called a protein dynamical transition. The underlying physical origin of this transition had been poorly understood in the past, although water was thought to be involved.
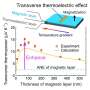
In their experiments, the researchers observed such a transition at -160 degrees Celsius (-256 Fahrenheit) -- a much lower temperature -- in a protein crystal when cryogenically cooled water confined in the crystal underwent unusual phase transitions.
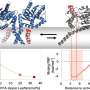
The protein crystal samples were treated by the researchers' own high-pressure cryo-cooling method. The samples were frozen to liquid nitrogen temperatures (-196 C, -320 F) in helium gas pressurized to 2,000 atmospheres. Previously, they had shown this method induces an unusual, high-density amorphous state of water, and, upon warming, it transforms to low-density amorphous ice.
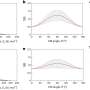
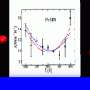
The protein dynamical transition was then probed at CHESS by temperature-controlled X-ray protein crystallography, which is typically used to measure the distribution of fluctuation states of a protein. The researchers observed that the fluctuation states suddenly increased as the water underwent the high-density to low-density transition.
The change in the state of water gave the proteins freedom to fluctuate and wiggle. This suggested that the protein dynamical transition was enabled by motional freedom provided by the surrounding water.
In the past, similar studies had been hampered by spontaneous crystallization of water into ice. The researchers' novel high-pressure method bypasses this problem to allow probing of protein dynamics and the relationship to the phase behavior of water at cryogenic temperatures. The results provide evidence that the physical origin of a protein dynamical transition is driven by water fluctuations, and they also provide insights into the unusual physical properties of supercooled water.
Provided by Cornell University