Engineers probe mechanics behind rapid-aging disease
Researchers at MIT and Carnegie Mellon University are using both civil engineering and bioengineering approaches to study the behavior of a protein associated with progeria, a rare disorder in children that causes extremely rapid aging and usually ends in death from cardiovascular disease before age 16. The disease is marked by the deletion of 50 amino acids near the end of the lamin A protein, which helps support a cell's nuclear membrane.
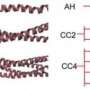
At MIT, the researchers used molecular modeling — which obeys the laws of physics at the molecular scale — to simulate the behavior of the protein's tail under stress in much the same way a traditional civil engineer might test the strength of a beam: by applying pressure. In this instance, they created exact replicas of healthy and mutated lamin A protein tails, pulling on them to see how they unraveled.
"The application of engineering mechanics to understand the process of rapid aging disease may seem odd, but it actually makes a lot of sense," says Markus Buehler, a professor in MIT's Department of Civil and Environmental Engineering who also studies structural proteins found in bone and collagen. In this new research, he worked with Kris Dahl, professor of biomedical engineering and chemical engineering at Carnegie Mellon, and graduate students Zhao Qin of MIT and Agnieszka Kalinowski of Carnegie Mellon. They published their findings in the September issue of the Journal of Structural Biology.
In its natural state, a protein — and its tail — exist in complex folded configurations that differ for each protein type. Many misfolded proteins are associated with diseases. In molecular simulations, Qin and Buehler found that the healthy lamin A protein tail unravels sequentially along its backbone strand, one amino acid at a time.
"It behaved much as if I pulled on a loose thread on my shirt cuff and watched it pull out stitch by stitch," said Qin.
By contrast, the mutant protein tail, when pulled, first breaks nearly in half, forming a large gap near the middle of its folded structure, then begins unfolding sequentially. The MIT scientists deduced that it takes an additional 70 kilocalories per mole (a unit of energy) to straighten the mutant tails, meaning the mutant protein is actually more stable than its healthy counterpart.
At Carnegie Mellon, Dahl and Kalinowski studied the same topic by subjecting lamin A protein tails to heat, which causes proteins to denature or unfold. In their lab, they observed the same pattern of unraveling in healthy and mutated proteins as the MIT engineers did in their atomistic simulation.
Qin then wrote a mathematical equation to convert the temperature differential seen in denaturing the mutant and healthy proteins (4.7 degrees Fahrenheit) to the unit of energy found in the atomistic simulations, finding that the increase in temperature very nearly matched the increase in energy. This agreement, the researchers say, validates the application of the civil engineering methodology to the study of the mutated protein in diseased cells.
The results, however, were counterintuitive to the civil engineers, who are accustomed to flawed materials being weaker — not stronger — than their unimpaired counterparts.
As a component of the cell's nucleoskeleton, lamin A plays an important role in defining the mechanical properties of a cell's nuclear membrane, which must remain flexible enough to easily withstand deformation. In previous work, Dahl had observed that nuclear membranes built from the mutated proteins become very stiff and brittle, which could explain the altered protein-DNA and protein-protein interactions observed in diseased cells.
"Our surprising finding is that the defective mutant structure is actually more stable and more densely packed than the healthy protein," said Buehler. "This is contrary to our intuition that a 'defective' structure is less stable and breaks more easily, which is what engineers would expect in building materials. However, the mechanics of proteins is governed by the principles of nanomechanics, which can be distinct from our conventional understanding of materials at the macro scale."
Provided by Massachusetts Institute of Technology