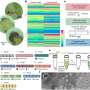
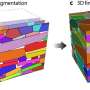
Typical stress response for algae influences photosynthetic productivity
Typical research on blue-green algae, or cyanobacteria, often focuses on pampered conditions that result in a fast, high yield, but such conditions bear little resemblance to what these bacteria face in nature. Scientists from Pacific Northwest National Laboratory (PNNL), Washington University in St. Louis, The Hebrew University of Jerusalem, and Texas Tech University conducted a large-scale analysis of an alga called Synechocystis 6803 under 33 environmental conditions.
The scientists discovered that, regardless of the type of stress or duration, the cyanobacterium immediately activates alternate pathways to acquire carbon and nitrogen, needed for survival. Analysis of these dynamic changes in the proteome provides insights into cellular adaptations to environmental perturbations.
By using this type of large-scale approach, scientists can gain insights into how these important bacteria function in nature. Cyanobacteria are responsible for nearly half of the photosynthesis necessary for sustaining life on earth. They've been studied by hundreds of laboratories over decades, yet many aspects of their physiology remain poorly understood. Better understanding of cyanobacteria will allow engineers to use them more effectively in the generation of renewable, carbon-neutral biofuels.
The team has discovered how these critical bacteria use many proteins and how they respond to varying natural conditions. In this study, they applied large-scale mass spectrometers toward a global proteomic analysis—a capability at EMSL, a Department of Energy scientific user facility at PNNL, to produce a high-quality dataset that covers 53 percent of the predicted proteome. This is the most comprehensive protein based functional and quantitative analysis for any photosynthetic organism to date.
More information: Wegener KM, et al. 2010. "Global Proteomics Reveal an Atypical Strategy for Carbon/Nitrogen Assimilation by a Cyanobacterium Under Diverse Environmental Perturbations." Molecular & Cellular Proteomics 9(12):2678-2689. DOI:10.1074/mcp.M110.000109
Provided by Pacific Northwest National Laboratory