Scientists devise broad new technique for screening proteins
A team led by scientists from The Scripps Research Institute has developed a powerful new method for detecting functional sites on proteins. The technique may have broad applications in basic research and drug development.
Described in an advance, online publication of Nature on November 17, 2010, the method enables scientists to take a sample of cells, locate the sites on their proteins that have a certain kind of biochemical reactivity, and measure the degree of that reactivity.
"It lets us find functional sites on proteins more efficiently than before, and that's going to be helpful not only for characterizing unknown proteins, but also for finding new sites of importance on already-characterized proteins," says the study's senior investigator, Benjamin F. Cravatt III, PhD, professor and chair of the Department of Chemical Physiology and member of the Skaggs Institute for Chemical Biology at Scripps Research in La Jolla, California.
The Hyper-Reactive Sites
Scientists already have techniques for identifying the sequence of amino acids that make up a given protein. But this sequence data doesn't tell them all they need to know. Initially translated from genetic material as a simple chain of amino acids, a protein thereafter typically folds into a complex, three-dimensional structure. The hard-to-predict details of this structure can strongly determine which sites on a protein are highly reactive.
Scientists can find these reactive sites with biochemistry studies, but traditionally such studies have required months or even years of work, even for a single protein. Partly for this reason, tens of thousands of proteins in humans and other species remain uncharacterized.
"What we've needed is a more efficient method to find and quantitatively analyze reactive sites," said Cravatt, "not just for one protein in a purified sample but for a large set of proteins in their natural setting, such as within a whole cell or tissue."
In the study reported in Nature, Cravatt's team, led by Research Associates Eranthie Weerapana, PhD, and Chu Wang, PhD, set out to develop just such a method. As a proof of principle, they targeted cysteine, one of the most reactive amino acids found on proteins. "A cysteine site on a protein often is responsible for enzymatic activity, or serves as an anchoring point for a chemical modification that regulates the protein's activity," says Wang.
A Dynamic Duo of Probes
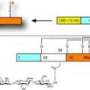
The approach the team developed involved the creation of special kind of cysteine-labeling chemical probe – in fact, two probes, which were chemically the same but differed very slightly in their mass so that they could later be distinguished. Typically, probes that label cysteines are used at a high enough concentration to mark all the cysteines in a given sample. In this case, the two probes were added to cellular proteins at a concentration about a hundred times lower than normal, so that for most cysteines, the labeling would be incomplete. Another novel element of the scheme was that the concentration of the probes was varied, so that one protein sample was treated with up to ten times higher probe than the other.
"Our hypothesis was that the hyper-reactive cysteines of interest would be fully labeled by these probes even at very low probe concentrations, and thus the low- and high-concentration probes would label these cysteines to the same degree," says Wang. "The cysteines with ordinary reactivity, by contrast, would be incompletely labeled at low probe concentrations and would show greater labeling as the probe concentration became higher."
The team first tested the technique on the proteins found within a human breast cancer cell line, and soon identified and located more than 800 cysteine sites on 522 proteins. For more than 90 percent of these cysteines, the low and high concentration probes showed correspondingly low and high levels of labeling, indicating that the cysteines had ordinary reactivity.
"But a small fraction of the cysteines showed a constant level of labeling for low and high concentration probes," said Wang, "indicating that they were hyper-reactive."
At first by examining select proteins from this group, and later by looking up all 522 labeled proteins in a database, the team found that the cysteines marked as hyper-reactive by their technique were highly enriched in known functional sites. The non-hyper-reactive cysteines, by contrast, were much less commonly listed as functional.
"By extrapolation, we can say that cysteines that haven't yet been officially characterized, but which show this hyper-reactivity in our assay, are likely to be functional," said Cravatt. To lend further support to this hypothesis, the team performed experiments, in collaboration with Scripps Research Assistant Professor Kerri Mowen, Ph.D, on an uncharacterized hyper-reactive cysteine in this list and showed that it played an important functional role in its parent proteins.
In a final set of experiments, collaborating investigator David Baker and colleagues at the University of Washington supplied a set of synthetic proteins that had been designed to work as enzymes. Weerapana and Wang and their team were able to predict, using their reactive-cysteine tagging technique, which of these proteins had the hoped-for enzymatic function.
"This is a relatively precise and straightforward method for screening designed proteins for functional properties," said Cravatt. "It could be very useful for creating new enzyme catalysts for basic research and industrial applications."
And cysteine is only one type of amino acid to which this basic technique could be applied. "All you would have to do, in principle, is change the reactive group on the probe, and instead of targeting cysteines, target lysines or serines or tyrosines, or some other amino acid," Cravatt said. "I think the approach will have broad utility in many areas of biology."
Provided by The Scripps Research Institute