This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Rapid approach to creating cyclic peptide opens the way for new antibiotics
A discovery made by scientists at King's College London could speed up efforts to produce new antibiotics in the fight against antimicrobial resistance.
In a paper published in the Journal of the American Chemical Society, scientists from the Department of Chemistry share a new, rapid method for making cyclic peptides—an important class of antibiotic molecules. The approach takes minutes rather than the hours or days it has normally taken, helping overcome a major challenge in antibiotic development.
Lead author Dr. Sarah Barry, from the Department of Chemistry at King's College London, said, "The global rise in antimicrobial resistant infections is putting the huge gains in modern medicine over the last century at risk. Everything from a cut in your finger to major surgery or cancer treatment requires the use of antibiotics. But when bacteria and viruses evolve to evade these medicines, these life-saving drugs lose their effectiveness.
"We urgently need to invest in antibiotic development—and we hope that breakthroughs like our new method at King's can inspire renewed efforts towards this goal."
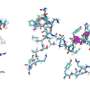
With antimicrobial resistance (AMR) a major and growing threat to public health around the world, new antibiotics are urgently required to fight deadly infectious diseases. Yet modifying or designing antibiotics to evade resistance is challenging due to the complexity of their molecular structure. Some are formed of cyclic peptides—strings of amino acids joined together in a circle—that are very difficult to replicate in the lab.
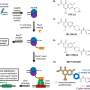
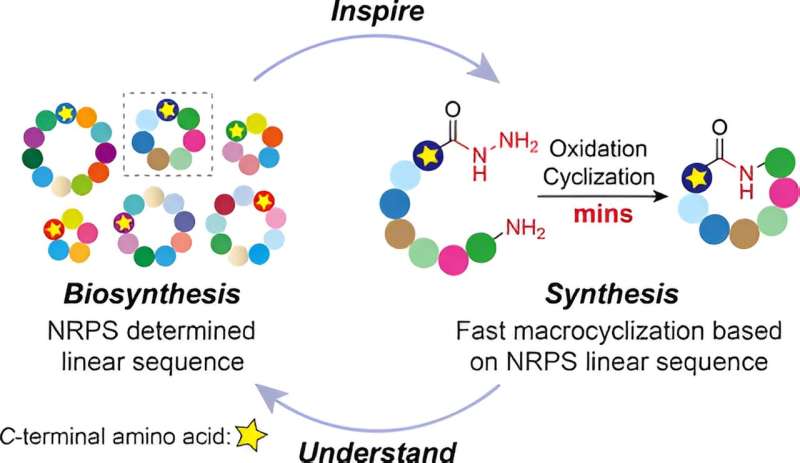
To understand how this process occurs in nature, the scientists studied a naturally produced cyclic peptide which has promising activity against the bacteria that causes TB—the world's deadliest infectious disease and a disease that is becoming more resistant. They found that when using the encoded sequence of amino acids from the natural peptide to make their synthetic peptides, it enabled their peptides to close into cycles incredibly quickly.
The team believes this simple approach can be used to make modified versions of the molecules used in antimicrobials, which can be used to develop new antibiotics.
Dr. Yaoyu Ding, Research Assistant, whose Ph.D. was based on the project, said, "We have made a key part of the chemistry involved in making these molecules far easier and quicker. We hope that this chemistry will enable researchers to make libraries of derivates to enable screening for new antimicrobials."
While their primary interest was in antimicrobials, the scientists have also successfully tested this process on a range of other peptides, opening the way for cyclic peptide discovery in a range of drugs, including anti-cancer agents.
More information: Yaoyu Ding et al, Rapid Peptide Cyclization Inspired by the Modular Logic of Nonribosomal Peptide Synthetases, Journal of the American Chemical Society (2024). DOI: 10.1021/jacs.4c04711
Journal information: Journal of the American Chemical Society
Provided by King's College London