This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
proofread
A multi-stream network for retrosynthesis prediction
Retrosynthesis aims to predict a set of reactants for producing given molecules, which plays a significant part in the biochemistry field, such as molecular pathway design and drug discovery. Most existing methods only benefit from one kind of information rather than further considering the diverse aspect of molecular information.
To address this issue, a research team conducted a study, now published in Frontiers of Computer Science.
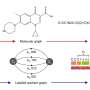
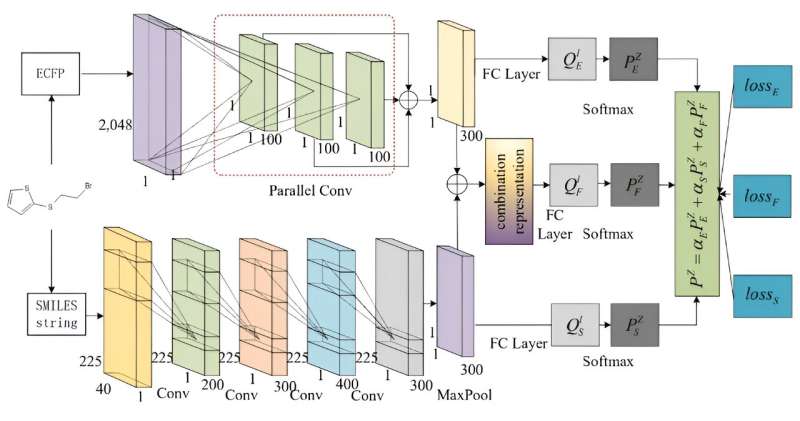
The team proposed a multi-stream network for retrosynthesis prediction by describing molecules from multiple perspectives by using their SMILES and ECFPs descriptors.
MSNR consists of three major modules: (i) The parallel CNN and the text-CNN that takes the ECFPs and SMILES after one-hot encoding as input to produce the deep features. (ii) The combination representation is devised by fusing the two kinds of deep features of ECFPs and SMILES, which provide an in-depth perspective of molecular representation. (iii) Three dense classifiers have been implemented to predict the probability of reactants for molecules, which leverages the deep features extracted by different streams as the molecular representation, respectively.
By fusing these multi-stream prediction results with varying weights, the model arrives at a final retrosynthesis prediction. Additionally, the model is trained using an overall loss function that is capable of exploiting the diverse information available from each type of deep feature.
More information: Qiang Zhang et al, A multi-stream network for retrosynthesis prediction, Frontiers of Computer Science (2023). DOI: 10.1007/s11704-023-3103-z
Provided by Frontiers Journals