This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
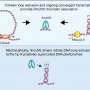
Machine learning method reveals chromosome locations in individual cell nucleus
Researchers from Carnegie Mellon University's School of Computer Science have made a significant advancement toward understanding how the human genome is organized inside a single cell. This knowledge is crucial for analyzing how DNA structure influences gene expression and disease processes.
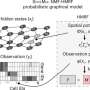
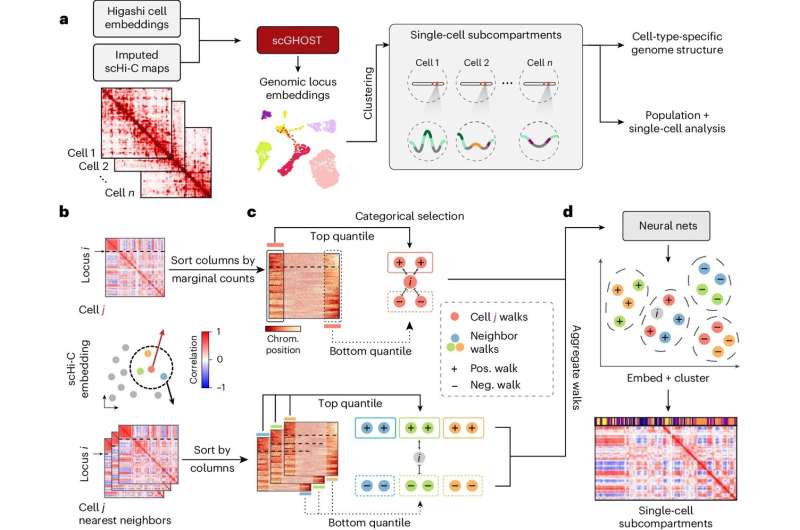
In a paper published by the journal Nature Methods, Ray and Stephanie Lane Professor of Computational Biology Jian Ma and former Ph.D. students Kyle Xiong and Ruochi Zhang introduce scGHOST, a machine learning method that detects subcompartments—a specific type of 3D genome feature in the cell nucleus—and connects them to gene expression patterns.
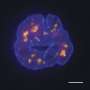
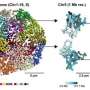
In human cells, chromosomes aren't arranged linearly but are folded into 3D structures. Researchers are particularly interested in 3D genome subcompartments because they reveal where chromosomes are located spatially inside the nucleus.
"One of the ultimate goals of single-cell biology is to elucidate the connections between cellular structure and function across a wide variety of biological contexts," Ma said. "In this case, we are exploring how chromosome organization within the nucleus correlates with gene expression."
While new technologies allow the study of these structures at the single-cell level, poor data quality can hinder precise understanding. scGHOST addresses this problem by using graph-based machine learning to enhance the data, making it easier to pinpoint and identify how chromosomes are spatially organized. scGHOST builds upon the Higashi method Ma's research group previously developed.
With the ability to accurately identify 3D genome subcompartments, scGHOST adds to the growing array of single-cell analysis tools scientists use to delineate the intricate molecular landscape of complex tissues, such as those in the brain. Ma anticipates that scGHOST could open new avenues to understanding gene regulation in health and disease.
More information: Kyle Xiong et al, scGHOST: identifying single-cell 3D genome subcompartments, Nature Methods (2024). DOI: 10.1038/s41592-024-02230-9
Journal information: Nature Methods
Provided by Carnegie Mellon University