This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Machine learning reveals sources of heterogeneity among cells in our bodies
A team of South Korean scientists led by Professor Kim Jae Kyoung of the Biomedical Mathematics Group within the Institute for Basic Science (IBS-BIMAG) discovered the secrets of cell variability in our bodies. The findings of this research are expected to have far-reaching effects, such as improvement in the efficacy of chemotherapy treatments, or the development of a new paradigm in the study of antibiotic-resistant bacteria.
Dr. Jo Hyeontae and Dr. Hong Hyukpyo participated as co-first authors in this research, which is published in the journal Patterns. The title of the paper is "Density Physics-informed Neural Networks Reveal Sources of Cell Heterogeneity in Signal Transduction."
The cells in our body have a signaling system that responds to various external stimuli such as antibiotics and osmotic pressure changes. This signaling system plays a critical role in the survival of cells as they interact with the external environment. However, even cells with same genetic information can respond differently to the same external stimuli, called cellular heterogeneity.
Cellular heterogeneity is a great research interest in medicine, as it is known to hinder the complete eradication of cancer cells by chemotherapeutic agents such as anticancer drugs. The sources of such heterogeneity and its relationship with the signaling system have remained a challenge, as intermediate processes of the signaling system are impossible to fully observe with current experimental technology.
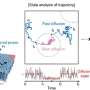
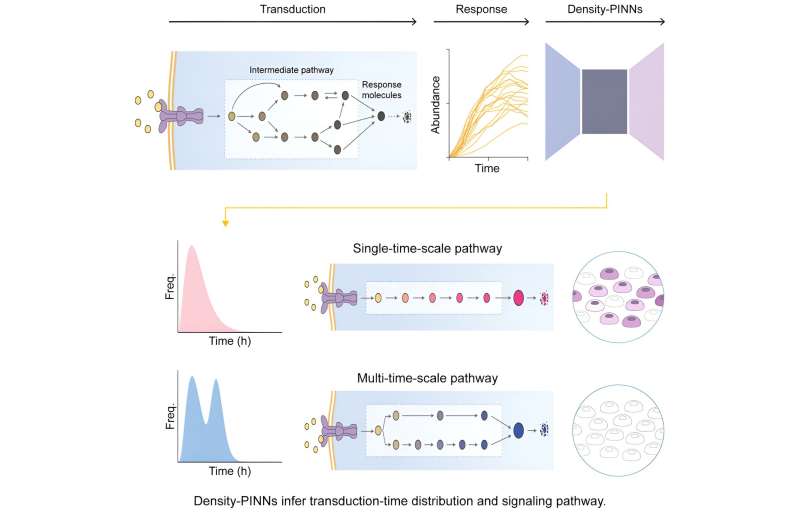
To reveal the sources of this heterogeneity, Professor Kim's research team developed a machine learning methodology using artificial neural network structures called Density Physics-informed neural networks (Density-PINNs). Density-PINNs use the observable time-series data of cells' responses to external stimuli to inversely estimate information about the signaling system.
By applying Density-PINNs to actual experimental data of antibiotic responses of bacterial cells (Escherichia coli), the research team found that a parallel structure of the signaling system can reduce heterogeneity among cells.
Professor Kim believes that this mathematical modeling and machine learning research will facilitate the enhancement of the understanding of cellular heterogeneity, which is crucial in cancer treatment. He expressed his hope that this achievement would lead to the development of improved cancer treatment strategies.
More information: Hyeontae Jo et al, Density physics-informed neural networks reveal sources of cell heterogeneity in signal transduction, Patterns (2023). DOI: 10.1016/j.patter.2023.100899
Journal information: Patterns
Provided by Institute for Basic Science