This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
trusted source
proofread
Novel insights into antibody aggregation expected to open up new avenues for research and therapeutic applications
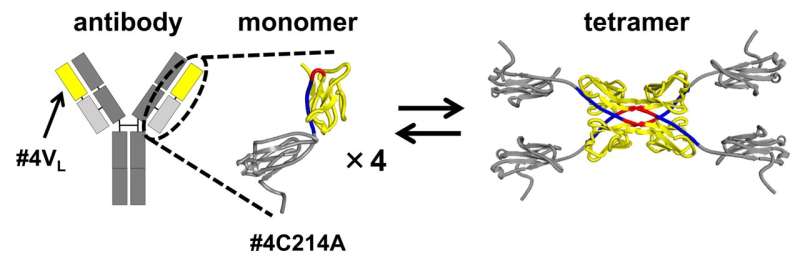
Antibodies (immunoglobulins) are Y-shaped proteins that recognize and neutralize specific pathogens. Their ability to target specific molecules or cells has made them promising candidates for future drug development. However, their light chains—parts of the antibody that contribute to recognizing and binding to specific antigens—misfold and aggregate, leading to amyloidosis, a condition that brings about complications and tissue dysfunction in the body.
In the context of drug development, antibody aggregation can compromise their capacity to bind to antigens and diminish their therapeutic potential. However, lack of detailed structural information on its aggregation is one of the factors hindering progress in the field.
As a result, ongoing efforts aim to provide detailed reports on aggregate structures and their formation mechanisms to advance antibody drug development.
In a study published in Nature Communications, a team of researchers from Japan, led by Shun Hirota from Nara Institute of Science and Technology (NAIST), has recently provided new insights into the structures formed during antibody aggregation through 3D domain swapping (3D-DS), a process where a specific region of a protein is exchanged between two or more molecules of the same protein.
The 3D-DS process has been observed in various proteins but not in antibody light chains until the present study.
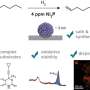
In their investigation, the researchers used a modified version of the antibody light chain. In this modified form, a cysteine (Cys) residue, which typically forms a disulfide bond with a heavy chain cysteine, was replaced with alanine (Ala). This alteration allowed the team to isolate and study the structures resulting from 3D-DS in the segment of the antibody contributing to antigen binding.
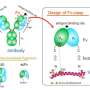
The 3D-DS of the antibody light chain involves the formation of dimers (structures consisting of two identical subunits) and tetramers (structures composed of two dimers with four identical subunits).
"Our study provides the first report on the atomic-level structure of the 3D-DS phenomenon in an antibody light chain's variable region," points out Hirota.
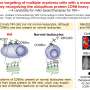
The size exclusion chromatography of the antibody light chain #4C214A revealed that the antibody exists as individual monomers and four-subunit tetramers. To determine the region where tetramers are formed, the researchers partitioned the antibody light chain into the variable region (the tip of the Y-shaped antibody) and the constant region (the middle part of the Y-shaped antibody).
They found that the variable region #4VL can switch between monomeric and tetrameric states.
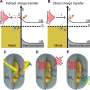
Further analysis using X-ray crystallography and thermodynamic simulations revealed that tetramer formation is driven by hydrophobic interactions occurring between two 3D-DS dimers.
Compared to monomers, the tetramers were found to have more rigid β-sheet structures, making them less flexible. The formation of the 3D-DS tetramer can help prevent protein aggregation by decreasing flexibility, potentially avoiding the formation of insoluble aggregates. On the other hand, 3D-DS may promote aggregation of antibodies.
Hirota concludes, "These findings not only clarify the domain-swapped structure of the antibody light chain but also contribute to controlling antibody quality and advancing the development of future molecular recognition agents and drugs."
More information: Structural and Thermodynamic Insights into Antibody Light Chain Tetramer Formation through 3D Domain Swapping, Nature Communications (2023). DOI: 10.1038/s41467-023-43443-4
Provided by Nara Institute of Science and Technology