This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Team discovers protein crucial for B cell differentiation and antibodies
A cell nucleus is a busy place. Cellular proteins twist and pull DNA, folding the genome into intricate 3D structures that support functioning of its coding parts.
This choreography is essential for cell development, and the exact steps vary wildly between cell types. Establishing proper communication between genes and far-away control switches at the right time in the right cell is not a small feat. In fact, very few proteins have the right combination of features to organize the genome into the right structures.
In a new study published in Cell, scientists at La Jolla Institute for Immunology (LJI) and Massachusetts General Hospital, Harvard Medical School, show how a protein called IKAROS helps "weave" the genome into the correct structure required for B cell differentiation and generation of a life-saving repertoire of antibodies.
"Without IKAROS, you cannot make a functioning B cell," says LJI Associate Professor Ferhat Ay, Ph.D., who co-led the new study.
Ay is an expert in genomic architecture. He investigates where genes are located—and how certain parts of the genome interact. His lab develops bioinformatics tools to create 3D maps of the genome, which is a key step in understanding the genetic underpinnings of immune cell development.
For the new research, Ay's laboratory partnered with Katia Georgopoulos, Ph.D., of Massachusetts General Hospital, Harvard Medical School, who pioneered previous studies showing that IKAROS is critical for immune cell development.
"IKAROS acts at the root of this development and enables differentiation of the hematopoietic stem cell into a diverse set of immune cells including B cells," says Georgopoulos. These earlier studies showed that IKAROS-loss-of function mutations caused lymphoid malignancies in animal models and were associated with poor prognosis in children and young adults with B cell precursor leukemias.
Could IKAROS serve as an early critical organizer of 3D genome organization needed for proper B cell development?
The Ay and Georgopoulos Labs joined forces to test this hypothesis by harnessing IKAROS mouse genetic models, B cell differentiation approaches, bioinformatics tools and 3D genomic maps to answer big questions about immune cell development.

Yeguang Hu, Ph.D., an instructor in the Georgopoulos Lab, performed technically demanding experiments to identify which parts of the genome touch each other with and without IKAROS at work. Daniela Salgado-Figueroa, a UC San Diego Bioinformatics Ph.D. student in the Ay Lab, led the analysis of the terabytes of complex data sets that were generated.
The researchers found that IKAROS solves a big problem in B cell development. B cells use receptors with two "arms" to detect pathogens. These arms have a "light chain" region and a "heavy chain" region. When a B cell recognizes a pathogen, it churns out antibodies with matching arms.
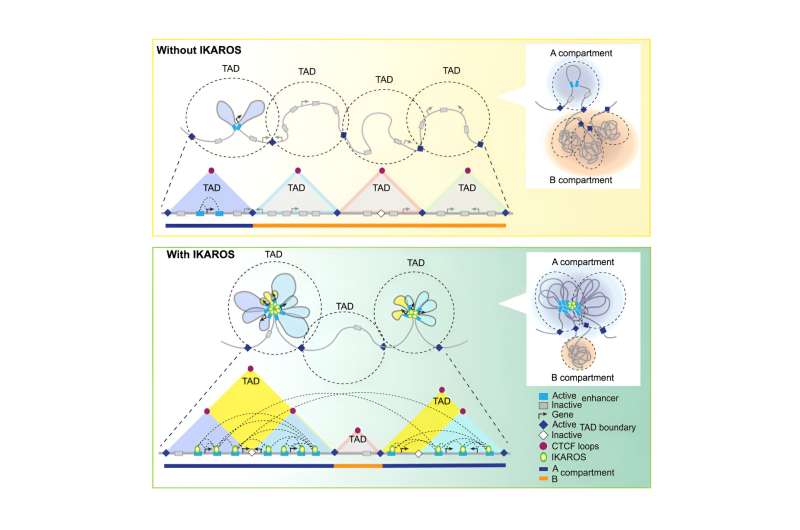
B cells assemble heavy chain regions fairly early in development. The Ay and Georgopoulos Labs found that assembling the light chain regions can be tricky because the genes encoding light chain development sit pretty far apart on the DNA. "This whole region needs to be rearranged in a proper 3D conformation," says Ay. "Something needs to bring them together."
Fortunately, IKAROS is on the scene to help with some genome gymnastics. The two labs found that IKAROS binds to specific parts of the genome and controls the formation of very useful loops using these sites as anchors. This looping brings far-away genes together with their control elements, leading to the activation and expression of the genes needed for proper B cell development and light chain rearrangement. Just as important, this folding of the genome keeps other control elements away from genes that should not be expressed in a B cell.
In a follow-up experiment, the researchers introduced IKAROS into human skin cells. They tested whether IKAROS could organize the genome in a cell type that did not produce any IKAROS of its own. And IKAROS prevailed. IKAROS, indeed, changed the 3D organization of a substantial part of the skin cell genome and led to formation of new loops, some of which resembled those from their B cell analysis.
The scientists emphasize that research into how chromatin is organized in the 3D space can help us understand how healthy cells develop—and how an improperly folded genome causes disease such as immunodeficiencies and cancer. Going forward, the researchers are interested in learning more about IKAROS disruption and disease development.
Additional authors of the study included Zhihong Zhang, Margaret Veselits, Sourya Bhattacharyya, Mariko Kashiwagi, Marcus R. Clark, and Bruce A. Morgan.
More information: Yeguang Hu et al, Lineage-specific 3D genome organization is assembled at multiple scales by IKAROS, Cell (2023). DOI: 10.1016/j.cell.2023.10.023
Journal information: Cell
Provided by La Jolla Institute for Immunology