This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Researchers capture high-resolution images of magnesium ions interacting with CRISPR gene-editing enzyme
The gene-editing technology known as CRISPR has led to revolutionary changes in agriculture, health research and more.
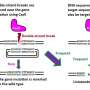
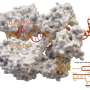
In research published in Nature Catalysis, scientists at Florida State University produced the first high-resolution, time-lapsed images showing magnesium ions interacting with the CRISPR-Cas9 enzyme while it cut strands of DNA, providing clear evidence that magnesium plays a role in both chemical bond breakage and near-simultaneous DNA cutting.
"If you are cutting genes, you don't want to have only one strand of DNA broken, because the cell can repair it easily without editing. You want both strands to be broken," said Hong Li, professor in the Department of Chemistry and Biochemistry and director of the Institute of Molecular Biophysics. "You need two cuts firing close together. Magnesium plays a role in that, and we saw exactly how that works."
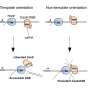
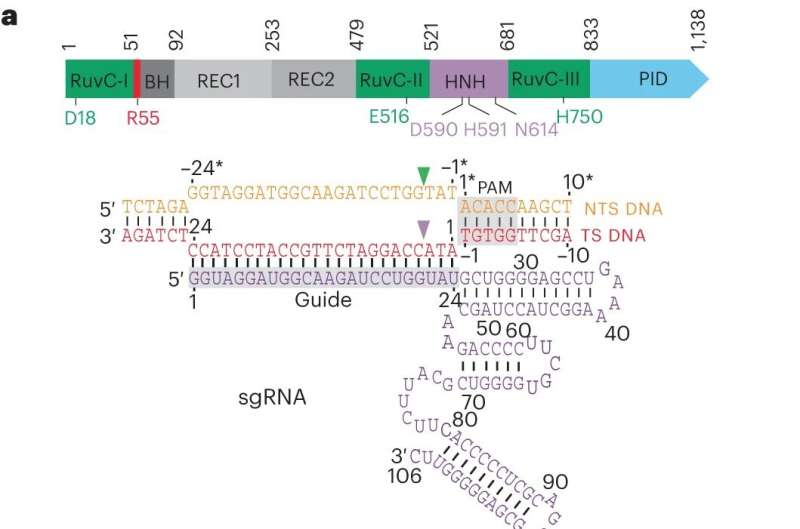
CRISPR-Cas9 is the most widely used tool for genetic manipulation. The technology uses a repurposed enzyme to bind to DNA, allowing alterations at specified locations in a genome.
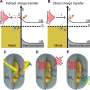
Scientists have known that magnesium plays a role in this process, but it was unclear exactly how, and no one had been able to capture time-lapsed images of the process up close. By leveraging a slower version of CRISPR-Cas9, this research showed that magnesium ions in the center of the catalysis reaction hold a key to the near-simultaneous cutting.
"I think a lot of times in science, even though you can infer something, you would like the proof," Li said. "For instance, with magnesium everybody knows you need it, but not seeing it in action, that's not complete science, right? You don't have the same level of understanding of how it functions."
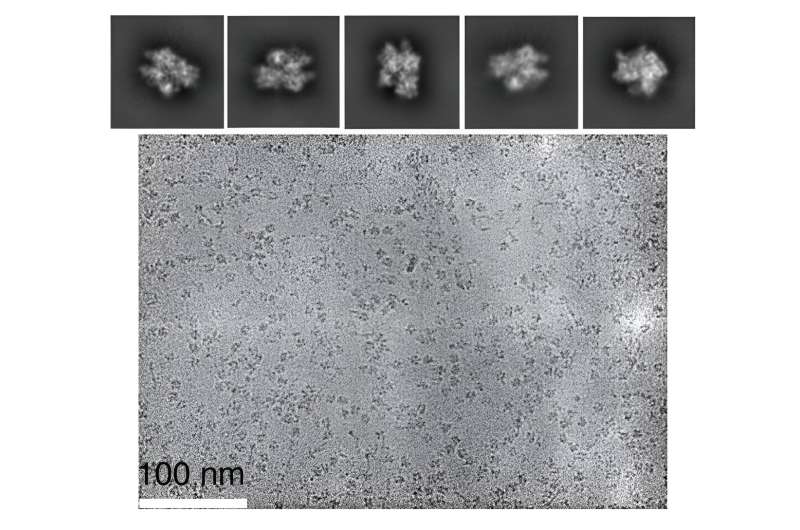

The researchers used the cryo-electron microscope at FSU's Biological Science Imaging Resource, which can produce images with near-atomic resolution, to observe metal ions and other atoms at work within the CRISPR-Cas9 enzyme. That allowed them to collect data that not only confirmed their earlier hypotheses but also led to the surprising discovery about how magnesium coordinates double-stranded breaks.
CRISPR made its debut in gene editing in 2013, and since then, scientists have worked to increase its dependability and expand its applicability to a variety of diverse organisms and cell types.
"By altering the active sites—the sets of 'scissors' that cut target and non-target DNA strands—we can sway the ability of Cas9 to use alternative metals for cutting," said doctoral candidate and paper co-author Mitchell Roth. "There's still a lot to explore with CRISPR."
Understanding how each element affects the enzyme's functioning gives scientists insight into what avenues for research might yield new knowledge and uses. Li and her team are planning further research to investigate how CRISPR-Cas9 can be retooled for other purposes.
Co-authors on this paper were former postdoctoral researchers Anuska Das and Jay Rai, doctoral candidate Yuerong Shu, undergraduate student Megan L. Medina and former undergraduate student Mackenzie R. Barakat, all of FSU.
More information: Anuska Das et al, Coupled catalytic states and the role of metal coordination in Cas9, Nature Catalysis (2023). DOI: 10.1038/s41929-023-01031-1
Journal information: Nature Catalysis
Provided by Florida State University