This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Revealing structural secrets of a key cancer protein
Scientists have breathed new life into the study of a protein with an outsized link to human cancers because of its dangerous mutations, using advanced research techniques to detect its hidden regions.
The Ras family of proteins are enzymes that set in motion the growth, division and differentiation of many types of cells, and their genes have been identified as the most frequently mutated cancer-related genes in humans. The subject of this study, the K-Ras protein, is linked to 75% of all Ras-associated cancers.
Researchers at The Ohio State University are the first to detect a section of this protein's structure that had previously been unobservable by standard lab tools, revealing features and interactions related to the protein's mutations that put cells into a state of perpetual division—a classic cancer characteristic.
"We know these mutations are a significant problem: They cause deaths," said senior study author Rafael Brüschweiler, Ohio Research Scholar and professor of chemistry and biochemistry at Ohio State. "We know that structural biology can provide unique insights into the mechanisms of those mutations and can stimulate the search for potential cures."
"We now have a more complete picture of what this protein does, which means we can start thinking about how to neutralize it once it's in its mutated form. Information in this sense is power, and this information is out there now so that we and other researchers can use it and start to hypothesize."
The study was published recently in the journal Nature Structural & Molecular Biology.
Despite existing knowledge about K-Ras and its key functional relationships with molecules related to cell health, the protein has been deemed "undruggable" because its configuration—both in normal and mutated forms—hides sites in its structure that would be most promising as therapeutic targets. Precision is required when designing such drugs—interfering with a protein in the wrong way could do more harm than the disease caused by a mutation.
"K-Ras is the holy grail of cancer research—probably one of the most studied biological molecules worldwide because it plays such a key role in many cancers," Brüschweiler said. "But it has also been a huge challenge."
Brüschweiler and colleagues reported in 2019 on a technique that enabled observation of proteins that move too slowly to be detected by standard nuclear magnetic resonance (NMR) spectroscopy. The team decided a year later to begin applying those findings to the hunt for K-Ras's secret hiding places.
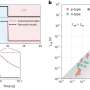
Standard NMR can follow a fast-acting protein but has trouble with a longer time scale of movement and interactions, and X-ray crystallography used to define protein structures does better with less movement and more time. Brüschweiler and colleagues could take into account both the dynamic nature of K-Ras as well as its interaction with the reactive ligand (GTP), first detecting faint signals from the hidden regions and then optimizing NMR experiments to strengthen those signals.
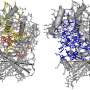
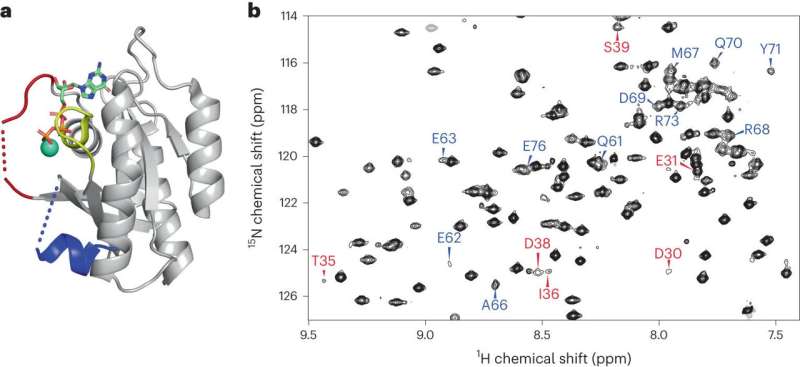
The study revealed two "switch" regions—tellingly, both located near a protein loop where the most dangerous mutations occur—in the K-Ras structure that had not been visible before. The team also established the complex structural dynamics behavior of the protein "backbone" that amplified additional features close to the switches. The backbone is essential to understanding a protein's structural properties—from there, characterizing amino acid side chains "is relatively straightforward," Brüschweiler said.
The experiments also added clarity to how the normal protein and its mutated forms differ: Under normal circumstances, K-Ras is more active when it is bound to the first of two partner molecules and maintains proper control of multiple cellular functions, including the return to an inactive state. When mutated, K-Ras gets stuck in the active phase and never takes a rest.
"We need active cells, but at some point they have to stop. Otherwise it's like never taking the foot off the accelerator in a car—at some point, you need to take your foot off because it's going too fast," he said. "That's the basic problem, that these mutations induce nonstop activity of the cell."
With the mutation-related switch regions now characterized, researchers have new drug targets to consider that could stifle the mutations without hampering K-Ras's essential cell functions.
"The switches and related areas where the switches interact are new candidates, which we now can monitor at unprecedented detail," Brüschweiler said. "This may not change the world overnight, but this is fundamentally new knowledge that has the potential to impact the health of human beings."
Brüschweiler has his own thoughts on what might come next, such as describing how existing drugs interact with the protein. Future work by his team and others will be supported by a new NMR instrument with a magnetic field of 1.2 gigahertz—which will be the most powerful NMR instrument in the United States—that has just arrived at Ohio State, where Brüschweiler is the principal investigator of the National Gateway Ultrahigh Field NMR Center.
More information: Alexandar L. Hansen et al, Excited-state observation of active K-Ras reveals differential structural dynamics of wild-type versus oncogenic G12D and G12C mutants, Nature Structural & Molecular Biology (2023). DOI: 10.1038/s41594-023-01070-z
Journal information: Nature Structural & Molecular Biology
Provided by The Ohio State University