Structural studies offer 'how-to' guide for designing cancer drugs
To design drugs that stall the growth of aggressive cancers, it helps to know the structures of the proteins that are revving the cancers' engines.
In a series of three papers published in Proceedings of the National Academy of Sciences, Scripps Research scientists have illuminated the three-dimensional structure of phosphoinositide 3-kinase alpha (PI3Kα), a protein often mutated in cancer cells. Moreover, the research team shed light on how that structure changes with the cancer-associated mutations, paving the way for drugs that could target only the mutated versions.
"We hope that these detailed structural findings lead to the discovery of drugs that affect cancer cells but not healthy cells," says senior author Peter Vogt, Ph.D., a professor in the Department of Molecular Medicine at Scripps Research. "That could potentially eliminate the side effects associated with current PI3Kα drugs."
PI3Kα plays a central role in cell survival and growth. In healthy cells, the protein is flipped on and off as needed. But in numerous types of cancer—including breast, colorectal, endometrial and brain—mutations in PI3Kα make it active all the time, encouraging the unchecked growth of the tumors.
Current drugs that aim to put the brakes on PI3Kα bind to a section of the protein that rarely changes between healthy and mutated versions; this means all the PI3Kα in the body is shut off. Because of that, these PI3Kα inhibitors carry a long list of side effects and toxicities.
"To solve this problem, you have to make inhibitors that only recognize the mutated versions of PI3Kα," says Vogt. "But to do that, you need structural information about what differentiates mutated, overactive PI3Kα from normal PI3Kα."
This is no easy feat: PI3Kα is a particularly flexible, "wiggly" protein, so it's difficult to get a single snapshot of its structure. Vogt's group, however, discovered that when PI3Kα was bound to one of the existing inhibitors, it became more stable.
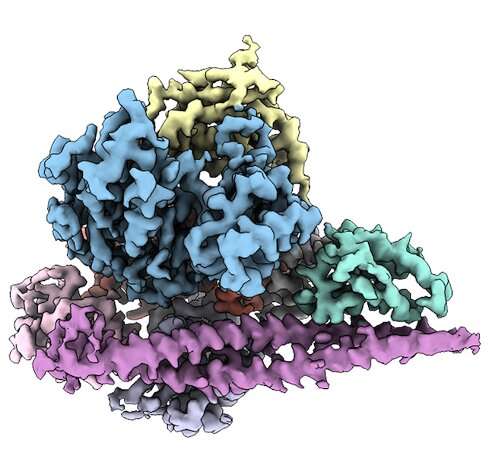
In PNAS papers published in November 2021 and September 2022, they used a type of imaging technique known as cryogenic electron microscopy (cryo-EM) to work out the three-dimensional structure of PI3Kα. With this knowledge, they first examined the structure of PI3Kα attached to the inhibitor. Then, to visualize the protein without the inhibitor, they used cross-linking molecules to attach different parts of PI3Kα to itself, stabilizing the most flexible parts of the protein.
More recently, the research team used the same cryo-EM toolbox to piece together the structure of two mutated versions of PI3Kα often found in cancer cells. That work, published last month in PNAS, showed how some segments of the mutated PI3Kα resemble the activated form of PI3Kα.
"There are quite dramatic structural changes," says Vogt. "And in the end, the changes essentially mimic the normal activated form of the protein, with the only difference being that it's always in this active structure."
The findings point toward ways to use drugs to shut off this always-on version of PI3Kα in cancer cells, without turning off healthy PI3Kα. The key, Vogt says, is that the drugs will need to bind to a different part of the PI3Kα protein than where the existing PI3Kα inhibitors bind—a part that varies structurally between the healthy and mutated versions of the protein.
His lab group is following up on this research with additional studies revealing how current drugs change the structure of PI3Kα.
More information: Xiao Liu et al, Cryo-EM structures of cancer-specific helical and kinase domain mutations of PI3Kα, Proceedings of the National Academy of Sciences (2022). DOI: 10.1073/pnas.2215621119
Journal information: Proceedings of the National Academy of Sciences
Provided by The Scripps Research Institute