This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Timing plant evolution with a fast-ticking epigenetic clock

Recent discoveries in the field of epigenetics, the study of inheritance of traits that occur without changing the DNA sequence, have shown that chronological age in mammals correlates with epigenetic changes that accumulate during the lifetime of an individual.
In humans, this observation has led to the development of epigenetic clocks, which are now extensively used as biomarkers of aging. While these clocks work accurately from birth until death, they are set back to zero in each new generation.
Now, an international team co-led by the University of Georgia, the GEOMAR Helmholtz Centre for Ocean Research Kiel and the Technical University of Munich, shows that epigenetic clocks not only exist in plants, but that these clocks keep ticking accurately over many generations. In a new study published in the journal Science, the team describes how this clock can tell time with a resolution from decades to centuries, an accuracy that cannot be achieved with traditional DNA mutation-based clocks.
The research sheds new light on microevolutionary questions that have been challenging to resolve, such as the timing of introduction of invasive species and the consequences of human activities since the emergence of modern industrialization.
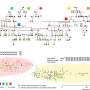
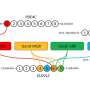
"Our first hint that an epigenetic clock exists in plants was revealed when we studied how DNA methylation, a chemical modification to DNA sequence underlying many epigenetic processes, varied across numerous branches in a 300-year-old poplar tree," said Frank Johannes, professor of plant epigenomics at the Technical University of Munich and co-author of the study. "We combined DNA methylation data with branch diameter and coring data to count tree rings, which reflects branch age. We were unable to core one branch, but we accurately estimated its age using only DNA methylation data, which provided the first clues there exists an epigenetic clock in plants."
The team's research showed experimentally that epigenetic clocks recapitulate known divergence times of intra-species phylogenetic or evolutionary trees in the self-fertilizing plant A. thaliana, a small plant in the mustard family, and the clonal seagrass Z. marina, which represent two major modes of plant reproduction.
"We further strengthened the existence of a plant epigenetic clock using a variety of experimental evolution populations of A. thaliana with known pedigrees," said Robert Schmitz, UGA Foundation Professor in Plant Sciences, Lars G. Ljungdahl Distinguished Investigator in the department of genetics, and co-author on the study.
These plants were grown by single-seed descent for up to 32 generations from wild type exposed to different environments or from natural strains from distinct geographical origins.
"Using DNA methylome data from hundreds of individuals from throughout these populations, we identified a subset of epimutations that are 'clock-like' and accurately estimated time of the pedigree," said Zhilin Zhang, doctoral student from the Technical University of Munich and co-lead author on the study along with Nan Yao, a doctoral student in the UGA Franklin College of Arts and Sciences department of genetics.
"We showed this epigenetic clock was more accurate at dating a recently diverged North American population of A. thaliana, approximately 140 years old, compared to a molecular clock using DNA mutations of the same individuals," Yao said.
"The proposed novel molecular clock will enable us to solve a long-standing riddle," said Thorsten Reusch, head of marine evolutionary ecology, GEOMAR Helmholtz Centre for Ocean Research Kiel. "Namely how old very large clones of fern, reed or seagrasses really are."
The study, "An evolutionary epigenetic clock in plants," was published Sept. 29.
More information: N. Yao et al, An evolutionary epigenetic clock in plants, Science (2023). DOI: 10.1126/science.adh9443
Journal information: Science
Provided by University of Georgia