This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Learning how to control HIV from African genomes
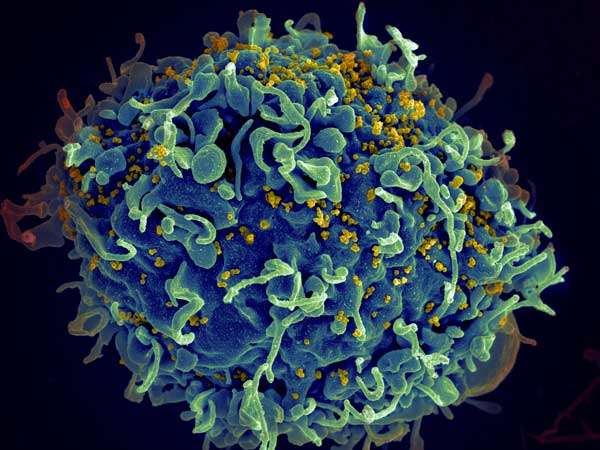
A study on almost four thousand people of African descent has identified a gene that acts as natural defense against HIV by limiting its replication in certain white blood cells. An international effort co-led by EPFL, Canada's National Microbiology Laboratory, and Imperial College London, it paves the way for new treatment strategies and underscores the importance of studying diverse ancestral populations to better address their specific medical needs and global health disparities.
"We searched for human genetic variation that associates with spontaneous control of HIV and identified a novel region in the genome that is only variable in populations of African ancestries," says Professor Jacques Fellay at EPFL's School of Life Sciences. "We used a combination of computational and experimental approaches to explore the biological mechanism behind the genetic association and provide evidence that the gene CHD1L acts to limit HIV replication in a subset of white blood cells."
HIV is still a problem
Despite significant advances in treatment and access to therapy, the human immunodeficiency virus remains a global health challenge with almost 40 million affected individuals, no vaccine and no cure. The virus attacks the person's immune cells (helper T cells, macrophages, and dendritic cells) damaging their ability to mount an immune response. Without treatment, the infected person grows more susceptible to opportunistic infections and cancer, and can develop acquired immunodeficiency syndrome, the well-known AIDS.
Although annual HIV infections have been declining because of widespread antiretroviral therapies, the trend has slowed substantially since 2005, and there are now alarming increases in the number of newly infected adults in some regions.
HIV and studies on the human genome
The way to therapies involves fundamental research, including studies into the relationship between the human genome and the progression of HIV infection, which can reveal possible therapeutic targets.
These Genome-Wide Association Studies, or GWAS, analyze the entire genome of a large number of individuals to identify genetic variants associated with a clinical outcome, such as the ability to naturally control viral replication.
Measuring HIV replication control: not enough in African populations
The degree of viral infection is measured by the virus' "setpoint viral load" (spVL), which refers to the relatively stable level of HIV replication in the body after the initial, acute phase of infection in untreated individuals.
A critical determinant of HIV infection progression and transmissibility, spVL is expressed as the number of viral copies per milliliter of plasma. The spVL of HIV varies widely in the infected population, depending on the ability of every individual's immune system to control viral replication without antiretroviral drugs.
Although there have been large studies of spVL control in populations of European descent, much less has been done in populations of African ancestries, which are still drastically underrepresented in human genomic studies. This is both a significant problem considering the disproportionate HIV burden in Africa and a missed opportunity given the high genome diversity among people of African descent, which fosters a high probability of genetic discoveries.
A key gene for resistance to HIV replication in people of African ancestries
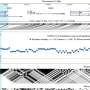
To address this disparity, a large international collaboration of scientists and clinicians has now performed large-scale GWAS using data from diverse populations of African ancestries. In total, the scientists analyzed the genomes from 3,879 individuals living with HIV-1. Using computational analysis and fine-mapping techniques, they identified a novel region in the genome that shows a strong association with spVL control.
The study was co-led by Jacques Fellay at EPFL, Paul McLaren at the Public Health Agency of Canada's National Microbiology Laboratory, and Manjinder Sandhu at Imperial College London. It is now published in Nature.
This region corresponds to a gene known as CHD1L (for "Chromodomain Helicase DNA Binding Protein 1 Like"), which encodes a protein that helps DNA unwind after it has been damaged, allowing it to be repaired. But in this study, the CHD1L gene showed genetic variation specific to populations of African ancestries, and that was linked to the spontaneous control of the most common and virulent type of HIV, called HIV-1.
Having identified CHD1L as a potential modulator of HIV-1 infection, the researchers explored the biological mechanism behind the genetic association and determined that CHD1L plays a role in limiting HIV replication in a subset of white blood cells.
The discovery of CHD1L's role in limiting HIV replication could lead to improved treatment options for infected individuals. "Our findings provide insights into potential therapeutic targets, which are needed to continue the fight against HIV-1," says Fellay. "In addition, our results underscore the importance of performing genomic studies in diverse ancestral populations to better address their specific medical needs and global health inequities."
More information: Jacques Fellay, Africa-specific human genetic variation near CHD1L associates with HIV-1 load, Nature (2023). DOI: 10.1038/s41586-023-06370-4. www.nature.com/articles/s41586-023-06370-4
Journal information: Nature
Provided by Ecole Polytechnique Federale de Lausanne