This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
proofread
Targeting resistance to a crucial reserve antibiotic

Colistin is a cationic cyclic peptide that disrupts bacterial cell membranes of Gram-negative bacteria. It is one of the few remaining antibiotics of last resort for use against infections with multidrug-resistant bacteria. Hence, the recent global detection of transferable mobile colistin resistance gene families in a wide range of multi-resistant Gram-negative bacteria isolated from all kinds of environments—clinical, veterinary, food-products and aquaculture—has sounded alarm bells.
Nevertheless, the success of mcr-1 as a transferable resistance factor remained puzzling, as its expression imposes a selective disadvantage on the growth properties of bacteria while imparting only moderate levels of resistance towards colistin.
Now, an international team led by scientists of the German Center for Infection Research (DZIF) uncovered why mcr-1—despite its drawbacks—is beneficial to bacteria. Their study suggests that acquiring mcr-1 induces a specific physiological state in bacteria that promotes resistance towards commonly encountered environmental stress conditions such as changes in acidity and antimicrobial peptides.
"We discovered that bacteria harboring mcr-1 trigger regulatory components of the bacterial envelope stress response, a system that senses fluctuations in nutrient availability and environmental changes. This in turn greatly increases MCR-1 production and promotes bacterial survival in low pH environments," says the paper's first author Dr. Renate Frantz from the Department of Medical Microbiology at Justus-Liebig University in Giessen, Germany.
The results suggest that integration of MCR-1-dependent resistance activity into the envelope stress response would support resistance of strains in environments that are stressful to bacterial cells, such as during passage through the gastrointestinal tract or when exposed to bile acids.
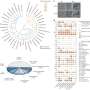
"Our analyses further showed, that production of the protein MCR-1 in bacteria grown under moderately acidic conditions leads to increased modification of lipid A, the anchor structure of lipopolysaccharides—a crucial component of the bacterial membrane required for colistin resistance," says Dr. Nicolas Gisch, joint first author from the Research Center Borstel, Leibniz Lung Center.
Based on the finding, that MCR-1 enzyme activity is greatly increased under acidic conditions, the research team developed a simple and easily reproducible assay to reliably determine MCR-1-dependent colistin resistance in bacterial isolates.
Production of the MCR-1 protein also induces expression of DegP, a protease in the bacteria's periplasm (the space between the inner and the outer membrane in Gram-negative bacteria), which cleaves MCR-1 at a specific site within a highly conserved region of the protein. "Modification of the cleavage site of MCR-1 has profound effects on both resistance activity and the triggering of the envelope stress response," adds co-lead author Dr. Konrad Gwozdzinski, former DZIF researcher and now a Senior Scientist at Mondelēz International.
These insights into the biomolecular basis of MCR-1-dependent resistance allowed the team to develop a general strategy that employs targeted activation of a protease to eliminate mcr-1-bearing plasmids from their bacterial hosts.
"Development of a simple diagnostic assay has important implications for ensuring the future use of the last-resort antibiotic colistin in clinical settings," comments Prof. Trinad Chakraborty, former Director of the Institute of Medical Microbiology at Justus Liebig University, who led the study. "Our data also allowed us to devise a novel approach to eliminate transferable colistin resistance in Gram-negative bacteria to counter its dissemination and spread in the environment."
The research is published in the journal Microbiology Spectrum.
More information: Renate Frantz et al, A Single Residue within the MCR-1 Protein Confers Anticipatory Resilience, Microbiology Spectrum (2023). DOI: 10.1128/spectrum.03592-22
Provided by Deutsches Zentrum für Infektionsforschung