This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
How skates learned to fly through water is revealed in their genome
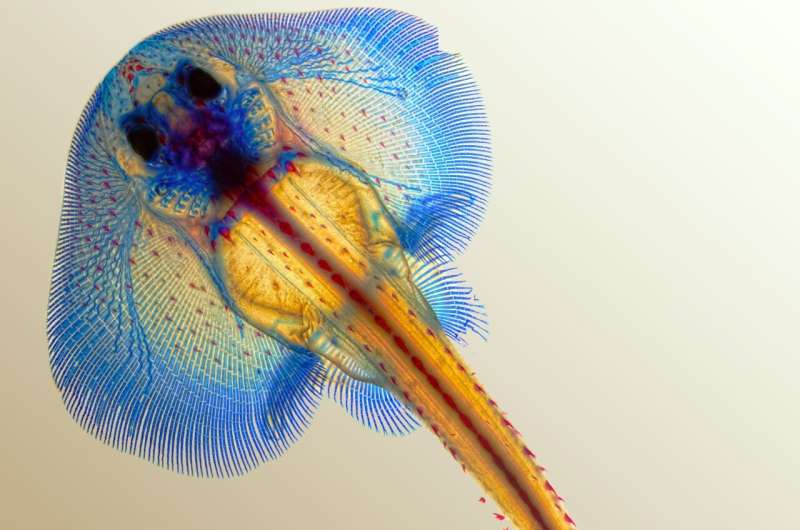
The little skate's dance on the ocean floor is graceful: Its massive frontal fins undulate as it skims beneath a layer of sand. With its mottled sand-colored camouflage, the animal is easy to miss.
Scientists at Max Delbrück Center in Berlin, the Andalusian Center for Developmental Biology (CABD) in Seville and other labs in the United States have discovered how the skate evolved these cape-like fins by peering into their DNA. They found that the key to the evolution of the skate fins lies not in the coding regions of its genome, but rather in the non-coding bits and the three-dimensional complexes that it folds into. These 3D structures are called topologically associated domains (TADs).
The international team describes in their article published in Nature that genomic changes that alter TADs can drive evolution. Until recently, genome evolution was mostly focused on studying variation at the DNA sequence level, but not in 3D genomic structures. "This is a new way of thinking about how genomes evolve," says Dr. Darío Lupiáñez, geneticist at the Max Delbrück Center and one of the lead authors of the study.
"Although we found that unique gene-expression patterns establish exceptionally wide skate fins a while ago, the underlying regulatory changes in the genome have previously remained unknown," says co-author Dr. Tetsuya Nakamura, developmental biologist at Rutgers University.
More than 450 million years ago, the genome of a primitive fish—the ancestor of all vertebrate animals—duplicated twice. The expansion in genetic material drove the rapid evolution of more than 60,000 vertebrates, including humans. One of our most distant vertebrate relatives are little skates (Leucoraja erinacea), which belong to a lineage of cartilaginous fishes that includes sharks and rays.
These distant cousins are ideal organisms to learn about the evolution of traits that made us human, such as paired appendages. "Skates are cartilaginous fishes called Chondrichthyans. They are considered more similar to ancestral vertebrates," says Dr. Christina Paliou, a developmental biologist at the CABD and one of the first authors. "We can compare the characteristics of skates with other species and determine what is novel and what is ancestral."
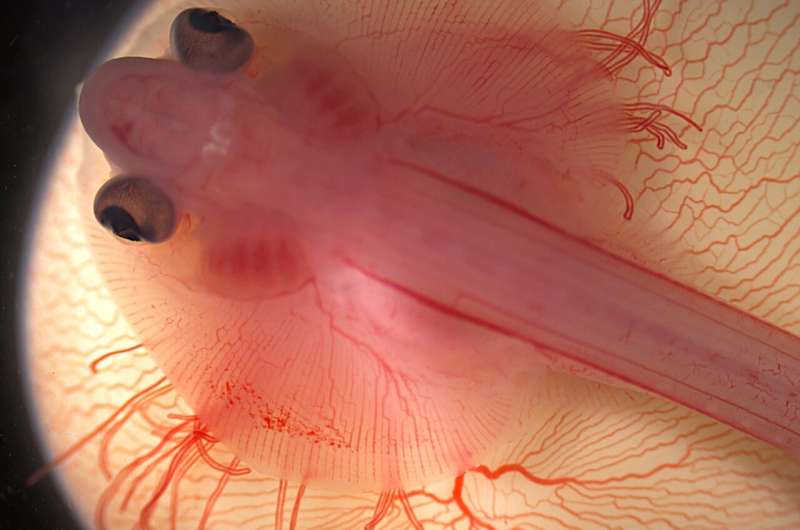
In 2017, the late Dr. José Luis Gómez-Skarmeta from the CABD, a founding figure in evolutionary genomics, brought together scientists from around the world to study skate evolution: laboratories with expertise in genome evolution such as the Ferdinand Marlétaz lab at University College London and Daniel Rokhsar lab at the University of California-Berkeley, in skate biology such as the Neil Shubin lab at University of Chicago, where Tetsuya Nakamura was then located (now at Rutgers) and in 3D gene regulation such as the Juan Tena at CABD, Darío Lupiáñez and Gómez-Skarmeta labs, as well as other collaborators.
Gómez-Skarmeta was interested in learning how genomes evolve structurally and functionally to promote the appearance of new traits. "To a great extent, evolution is the history of changing the regulation of gene expression during development," he said in 2018.
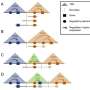
It was an exciting time for evolutionary genomics. Genome sequencing technologies had significantly improved and scientists could gain novel insights into how DNA, which stretches a couple of meters end-to-end, is folded into a 0.002-inch-diameter cell nucleus. "The packaging of DNA in the nucleus is far from random," says Lupiáñez. The DNA folds into 3D structures called TADs, which contain genes and their regulatory sequences. These 3D structures ensure that the appropriate genes are switched on and off at the right time, in the right cells.
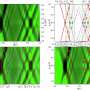
Dr. Rafael Acemel, a geneticist at the Max Delbrück Center and one of the first authors, performed experiments using the Hi-C technology, to elucidate the 3D structure of the TADs. But interpreting the results was challenging at first as the scientists needed the complete skate genome as a reference point. "At the time, the reference consisted of thousands of small unordered pieces of DNA sequence, so that did not help," Acamel says.
To overcome this difficulty, the scientists used long-read sequencing technology, together with Hi-C data, to assemble the pieces of the DNA like a puzzle and assign the unordered sequences to skate chromosomes. With the new reference, assembling the 3D structure of the TADs using Hi-C became trivial.
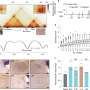
They compared this improved skate genome with genomes of the closest relatives, sharks, to identify any TADs altered during skate evolution. These altered TADs included genes of the Wnt/PCP pathway, which is important for the development of fins. There was also a skate-specific variation in a non-coding sequence near the Hox genes, which also regulate fin development. "This specific sequence can activate several Hox genes in the front part of the fins, which does not happen in other fish or four-legged animals," says Paliou. Subsequently, the scientists performed functional experiments that confirmed these molecular changes helped the skates evolve their unique fins.

TADs drive evolution
Earlier research has shown that changes in TADs can affect the expression of genes and cause diseases in humans. In this study, scientists show a role for TADs in driving evolution that has been previously noted for moles, too.
After the primitive fish ancestor duplicated its genome, many unused and redundant parts were subsequently lost. "It was not only the genes that disappeared, but also the associated regulatory elements and the TADs they are contained in," Lupiáñez says. "I think it's an exciting finding as it suggests that the 3D structure of the genome has an influence on its evolution."
TADs are important for gene regulation, 40% of them are conserved in all vertebrates, Acemel says. "However, 60% of TADs have evolved in some way or another. What were the consequences of these changes for species evolution? I think that we are just scratching the surface of this exciting phenomenon," Acemel says.
This mechanism of evolution constrained by TADs could be prevalent in nature. "We suspect that these mechanisms might explain many other interesting phenotypes that we observe in nature," Lupiáñez says. "By adding these new layers of gene expression, gene regulation, and 3D chromatin organization, the field of evolutionary genomics is entering into a new era of discovery."
More information: Daniel Rokhsar, The little skate genome and the evolutionary emergence of wing-like fins, Nature (2023). DOI: 10.1038/s41586-023-05868-1. www.nature.com/articles/s41586-023-05868-1
Journal information: Nature
Provided by Max Delbrück Center for Molecular Medicine