This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
How Polycomb repressive deubiquitinase specifically removes H2AK119 ubiquitination on nucleosome
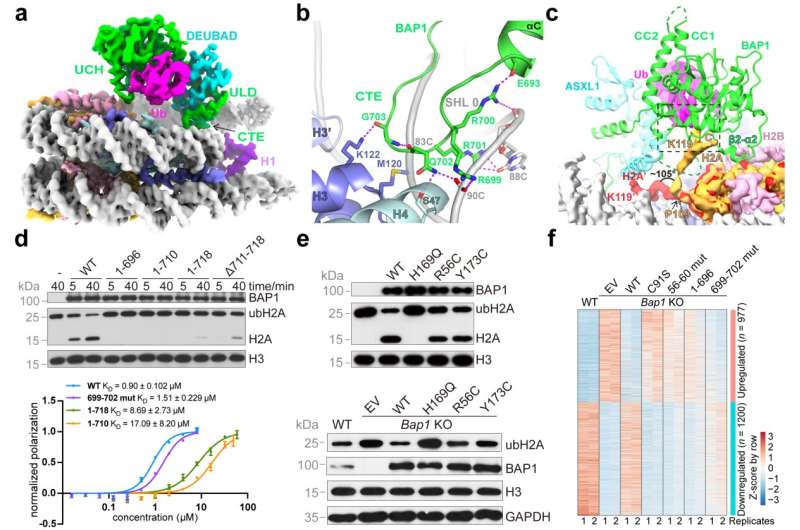
Chinese scientists reported the first high-resolution cryo-EM structure of human Polycomb repressive deubiquitinase (PR-DUB) complex bound to the chromatosome particle with mono-ubiquitinated H2AK119 (H2AK119ub1), revealing the mechanism by which PR-DUB specifically removes H2AK119ub1 in the context of nucleosome or chromatosome.
This study, led by Prof. Xu Ruiming and Zhu Bing's team from the Institute of Biophysics (IBP) of the Chinese Academy of Sciences (CAS), has been published in Nature.
Human PR-DUB complex is composed of ubiquitin C-terminal hydrolase BAP1 (BRCA1 associated protein 1) and one of the three Additional Sex comb Like (ASXL) proteins ASXL1/2/3. BAP1 is also a tumor suppressor that is frequently mutated in many cancers, and ASXL protein mutations are associated with varieties of leukemia.
Although the crystal structure of Drosophila PR-DUB complex has been previously reported, the solved structure lacked the ubiquitinated nucleosome substrate. This deficiency has severely limited the understanding of the molecular mechanism by which PR-DUB specifically removes H2AK119ub1 in the context of the nucleosome.
"Therefore, we determined the cryo-EM structure of PR-DUB in complex with the chromatosome, and found that amino acid residues 699–706 located on the positively charged BAP1 C-terminal extension (CTE) form a finger-like structure," said Xu. "It interacts with histone H3-H4 and DNA next to the nucleosome dyad."
Researchers located a highly conserved sequence motif, RRSRR, spanning residues 56–60 in the vicinity of the acidic patch of the nucleosome, in the catalytic domain of BAP1. This bidentate binding mode stabilizes the interaction between BAP1 and the nucleosome.
The ASXL1 subunit forms a complete ubiquitin binding pocket together with BAP1. Additionally, the interaction between ASXL1 and BAP1 directs the BAP1 CTE for correct nucleosome binding.
In this binding mode, the active site of BAP1 is located far away from the conventional H2AK119 position on the nucleosome, and the H2A C-terminal tail undergoes a large conformational change, placing the H2AK119-ubiquitin isopeptide bond in the catalytic active site for hydrolysis. The observed structural feature clearly accounted for the nucleosomal H2AK119 specificity of PR-DUB.
In addition, the researchers validated the structural observations by mutagenesis and in vitro and in vivo deubiquitinase assays of key residues at the interface between PR-DUB and nucleosome, and clarified the role of ASXL1 truncation mutants in the regulation of BAP1 enzymatic activity.
This study unveiled the particular nucleosome binding mode and the dramatic conformational change of ubiquitinated C-terminal tail of H2A in the nucleosome, revealing the structural basis for PR-DUB's specificity for nucleosomal H2AK119ub1.
"This revelation significantly expanded the understanding of the molecular mechanism governing deubiquitination of nucleosomal H2A, and the knowledge should also benefit the understanding of tumorigenesis effects of BAP1 and ASXL mutations and related drug development," said Prof. Xu.
More information: Weiran Ge et al, Basis of the H2AK119 specificity of the Polycomb repressive deubiquitinase, Nature (2023). DOI: 10.1038/s41586-023-05841-y
Journal information: Nature
Provided by Chinese Academy of Sciences