This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Discovery could lead to new fungicides to protect rice crops
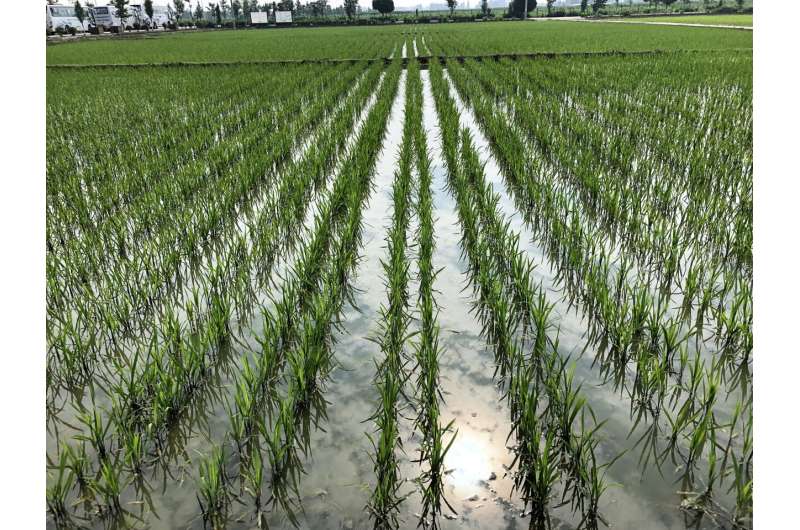

A fungus that plagues rice crops worldwide gains entry to plant cells in a way that leaves it vulnerable to simple chemical blockers, a discovery that could lead to new fungicides to reduce the substantial annual losses of rice and other valuable cereals.
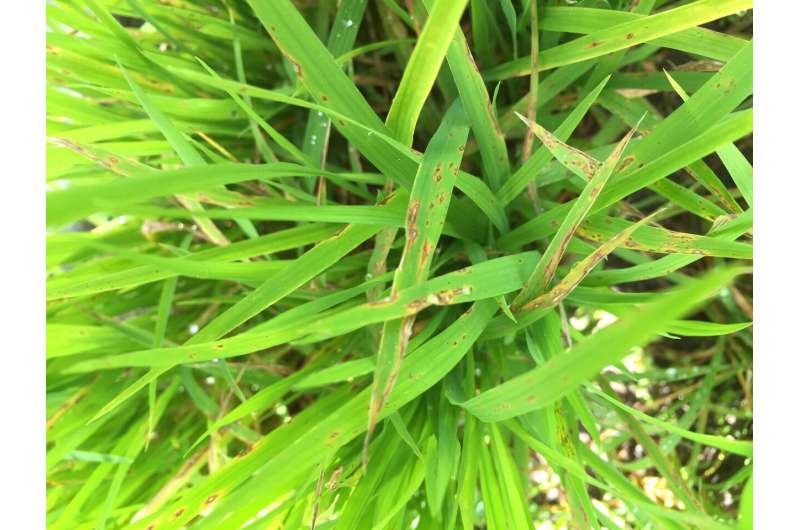
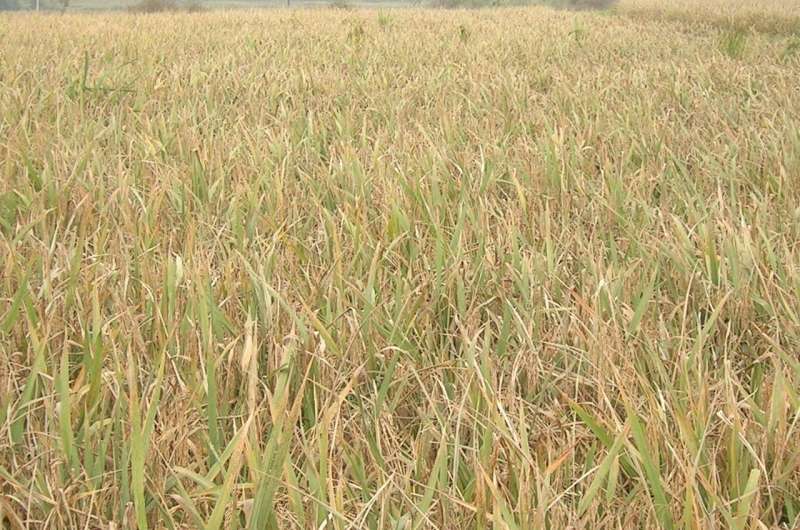
Each year, blast disease, caused by the fungal pathogen Magnaporthe oryzae, attacks and kills plants that represent between 10% and 35% of the global rice crop, depending on weather conditions.
University of California, Berkeley biochemists led by Michael Marletta, professor of chemistry and of molecular and cell biology, have discovered that the fungus secretes an enzyme that punches holes in the tough outer layer of rice leaves. Once inside, the fungus rapidly grows and inevitably kills the plant.
In a paper published this week in the journal Proceedings of the National Academy of Sciences, Marletta and his colleagues describe the structure of the enzyme and how it works to help the fungus invade plants. Because the enzyme is secreted onto the surface of the rice leaf, a simple spray could be effective in destroying the enzyme's ability to digest the wall of the plant. The scientists are now screening chemicals to find ones that block the enzyme.
"The estimates are that if you could knock out this fungus, you could feed 60 million more people in the world," said Marletta, the Choh Hao and Annie Li Chair in the Molecular Biology of Diseases at UC Berkeley. "This enzyme is a unique target. Our hope here is that we'll screen to find some unique chemicals and spin out a company to develop inhibitors for this enzyme."
This target is one of a family of enzymes called polysaccharide monooxygenases (PMO) that Marletta and his UC Berkeley colleagues discovered a little over 10 years ago in another more widespread fungus, Neurospora. Polysaccharides are sugar polymers that include starch as well as the tough fibers that make plants sturdy, including cellulose and lignin. The PMO enzyme breaks cellulose into smaller pieces, making the polysaccharide susceptible to other enzymes, such as cellulases, and speeding up the breakdown of plant fibers.
"There is an urgent need for more sustainable control strategies for rice blast disease, particularly in South Asia and sub-Saharan Africa," said Nicholas Talbot, who is Marletta's colleague and co-author, a plant disease expert and executive director of The Sainsbury Laboratory in Norwich in the United Kingdom. "Given the importance of the polysaccharide monooxygenase to plant infection, it may be a valuable target for developing new chemistries that could be applied at much lower doses than existing fungicides and with less potential environmental impact. It might also be a target for completely chemical-free approaches, too, such as gene silencing."

Marletta and UC Berkeley Ph.D students Will Beeson and Chris Phillips were originally interested in these enzymes because they degrade plant cellulose much more quickly than other previously described enzymes and thus had potential to turn biomass into sugar polymers that can be fermented more readily into biofuels. Fungi use PMOs to provide a source of food.
He and UC Berkeley colleagues subsequently found hints that some fungal PMOs may do more than merely turn cellulose into food. These PMOs were turned on in the early stages of infection, implying that they're important in the infection process rather than providing food.
That's what Marletta, Talbot and their colleagues found. Led by postdoctoral fellow Alejandra Martinez-D'Alto, the UC Berkeley scientists biochemically characterized this unique PMO, called MoPMO9A, while Talbot and UC Berkeley postdoctoral fellow Xia Yan showed that knocking out the enzyme reduced infection in rice plants.
Marletta and his UC Berkeley colleagues have found similar PMOs in fungi that attack grapes, tomatoes, lettuce and other major crops, which means the new findings may have broad application against plant fungal diseases.
"It isn't just rice that small molecule inhibitors could be used against. They could be widely used against a variety of different crop pathogens," Marletta said. "I think the future for this, in terms of drug development for plant pathogens, is pretty exciting, which is why we are going to pursue both the fundamental science of it, like we always do, and try to put together pieces to spin it out as a company."
Biofuels lead way to attacking fungal pathogen
Marletta specializes in identifying and studying new and unusual enzymes in human cells. But 10 years ago, when people got excited about biofuels as a way to address climate change, he was awarded a grant from UC Berkeley's Energy Biosciences Institute to search for enzymes in other life forms that digest plant cellulose faster than the enzymes known at the time. The goal was to turn tough cellulose fibers into short-chain polysaccharides that yeast could ferment into fuel.

"I said to two of my first-year graduate students, Chris Phillips and Will Beeson, "You know, there's got to be organisms out there that eat cellulose fast,'" Marletta said. "Those are the ones we want to find, because we know the enzymes that eat it slow, and they're not particularly useful in a biotechnology sense because they're slow."
Phillips and Beeson succeeded in finding fast-acting enzymes in a common fungus, Neurospora, which is among the first fungi to attack dead trees after a fire and does a quick job of digesting wood for nutrients. They isolated the enzyme responsible, the first known PMO, and described how it worked. Since then, Marletta's students have identified 16,000 varieties of PMO, most in fungi, but some in wood-eating bacteria. To date, these have had some success in speeding the production of biofuels as part of a cocktail of other enzymes, though they haven't made biofuels competitive with other fuels.
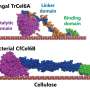
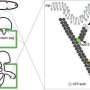
But Marletta was intrigued by a small subset of these 16,000 varieties that seemed to do more than provide nutrition for fungi. MoPMO9A, in particular, had an amino acid segment that binds to chitin, a polysaccharide that forms the outer coat of fungi, but is not found in rice. And though all PMOs are secreted, MoPMO9A was secreted during the infectious cycle of the fungus.
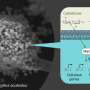
Studies subsequently showed that Magnaporthe concentrates MoPMO9A in a pressurized infection cell, called the appressorium, from which it is secreted onto the plant, with one portion of the enzyme binding to the outside of the fungus. The other end of the enzyme has a copper atom embedded in its center. When the fungus slaps the loose end of the enzyme onto the rice leaf, the copper atom catalyzes a reaction with oxygen to break cellulose fibers, helping the fungus breach the leaf surface and invade the entire leaf.
"We were curious: 'Hey, why does this enzyme have a chitin-binding domain if it's supposed to be working on cellulose?'" said Marletta. "And that's when we thought, "Well, maybe it's secreted, but it sticks to the fungus. That way, when the fungus is sitting on the plant, it can have between it and the leaf the catalytic domain to punch the hole into the leaf.'"
That proved to be the case. Marletta and Talbot are now testing other pathogens that produce PMOs to see if they use the same trick to enter and infect leaves. If so—Marletta is confident that they do—it opens avenues to attack them with a spray-on fungicide, as well.
"The only place you find PMOs like this is in plant pathogens that have to gain access to their host. So, they're almost certainly going to be working the same way," Marletta said. "I think the scope of work to develop inhibitors to this particular PMO is going to be well beyond rice, even though that itself is pretty important. We're going to be able to use them in other important crop plants."
More information: Martinez-D'Alto, Alejandra et al, Characterization of a unique polysaccharide monooxygenase from the plant pathogen Magnaporthe oryzae, Proceedings of the National Academy of Sciences (2023). DOI: 10.1073/pnas.2215426120. doi.org/10.1073/pnas.2215426120
Journal information: Proceedings of the National Academy of Sciences
Provided by University of California - Berkeley