Genome studies uncover a new branch in fungal evolution

About 600 seemingly disparate fungi that never quite found a fit along the fungal family tree have been shown to have a common ancestor, according to a University of Alberta-led research team that used genome sequencing to give these peculiar creatures their own classification home.
"They don't have any particular feature that you can see with the naked eye where you can say they belong to the same group. But when you go to the genome, suddenly this emerges," says Toby Spribille, principal investigator on the project and associate professor in the Department of Biological Sciences.
"I like to think of these as the platypus and echidna of the fungal world."
Spribille, Canada Research Chair in Symbiosis, is referring to Australia's famed Linnaean classification system-defying monotremes—which produce milk and have nipples, but lay eggs—that were the source of debate as to whether they were even real.
"Though nobody thought our fungi were fake, it's similar because they all look totally different."
Using DNA-based dating techniques, the team found that this new class of fungi, called Lichinomycetes, descended from a single origin 300 million years ago, or 240 million years before the extinction of dinosaurs.
David Díaz-Escandón, who performed the research as part of his Ph.D. thesis, explains that these "oddball" fungi were previously sprinkled across seven different classes—a high-level grouping that in animals would be equivalent to the groups called mammals or reptiles.
Working with a team of researchers from seven countries to get material from the fungi, he sequenced 30 genomes and found that all classes but one descended from a single origin.
"They were classified, but they were classified into such different parts of the fungal side of the tree of life that people never suspected they were related to each other," says Díaz-Escandón.
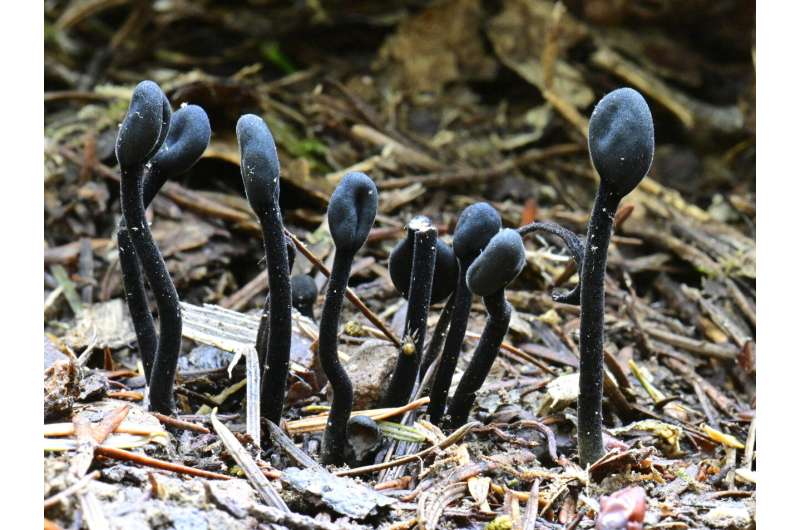
These fungi include forms as varied as earth tongues—eerie tongue-shaped fungi that shoot up vertically out of the ground—beetle gut microbes, and a fungus found in tree sap in northern Alberta. They also include some unusual lichens that survive in extreme habitats such as South America's Atacama Desert, the driest non-polar desert in the world.
"What is really fascinating is that despite these fungi looking so different, they have a lot in common at the level of their genomes," says Spribille. "Nobody saw this coming."
Based on their genomes, which are small compared with those of other fungi, the team predicts that this group of fungi depend on other organisms for life.
"Their small genomes mean this class of fungi have lost much of their ability to integrate some complex carbohydrates," said Spribille. "When we go back to look at each of these fungi, suddenly we see all of them are in a kind of symbiosis."
He notes the new research will be important to the broader study of fungal evolution, specifically how fungi inherit important biotechnological features such as enzymes that break down plant matter.
The new group also could be a source of new information about past fungal extinctions.
"We think it's likely that the diversity we see today is just the tip of the iceberg that survived. And we don't have that many examples of this kind of thing in fungi."
The research appears online in the journal Current Biology.
More information: Toby Spribille, Genome-level analyses resolve an ancient lineage of symbiotic ascomycetes, Current Biology (2022). DOI: 10.1016/j.cub.2022.11.014
Journal information: Current Biology
Provided by University of Alberta