Genomes of wild tomatoes offer valuable resource for tomato improvement
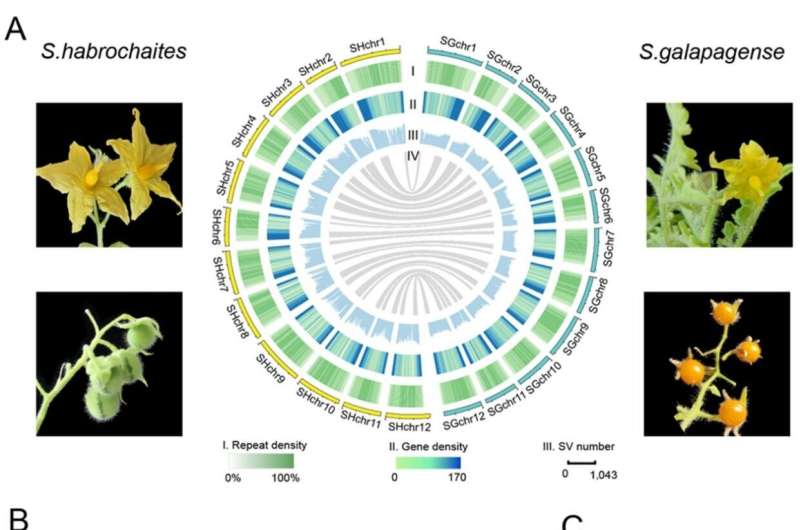
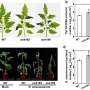
Wild tomato relatives Solanum habrochaites and S. galapagense are important germplasm donors in modern tomato breeding. Compared to the cultivated tomato, they display many desirable traits, particularly high tolerance to biotic and abiotic stresses. However, the lack of high-quality genomes has limited the study of the genetics and molecular mechanisms underlying these traits.
Researchers from the Wuhan Botanical Garden of the Chinese Academy of Sciences, together with collaborators from Cornell University and Zhejiang University, have generated the chromosome-scale reference genomes of the two species by combining PacBio HiFi reads and Hi-C sequencing data.
A phylogenetic tree constructed for S. habrochaites, S. galapagense and eight other Solanaceae species using 3,011 single-copy orthologous genes revealed that S. habrochaites was close to S. pennellii and S. galapagense appeared close to S. lycopersicum. Comparative analysis revealed 734 and 308 species-specific genes in S. habrochaites and S. galapagense, respectively, which might confer stress-tolerant trait of the two species.
A total of 336,319 structural variants (SVs) between S. habrochaites and S. lycopersicum and 98,443 SVs between S. galapagense and S. lycopersicum were identified, respectively. The inserted and expanded regions in the genomes of S. habrochaites and S. galapagense were found contributing to their tolerance to stresses like cold and pathogens, and unique terpene biosynthesis.
Terpene synthase genes (TPSs) were comprehensively characterized, an expansion of TPS-a subfamily was observed in S. habrochaites, which probably lead to its potentially diverse or unique sesquiterpene synthesis. A genome-wide identification of resistance gene analogs (RGAs) in cultivated tomato and four wild tomato relatives including S. habrochaites and S. galapagense, revealed 919 wild tomato-specific RGAs.
These results provide new genetic resources for terpenoids synthesis and disease resistance breeding in tomato.
This study has been published in Horticulture Research and is titled "Chromosome-scale genome assemblies of wild tomato relatives Solanum habrochaites and S. galapagense reveal structural variants associated with stress tolerance and terpene biosynthesis."
More information: Xiaofen Yu et al, Chromosome-scale genome assemblies of wild tomato relatives Solanum habrochaites and Solanum galapagense reveal structural variants associated with stress tolerance and terpene biosynthesis, Horticulture Research (2022). DOI: 10.1093/hr/uhac139
Provided by Chinese Academy of Sciences