How a protein breaks free to cause deadly cancers
Better treatments for some of cancer's deadliest forms could be closer due to a University of California, Irvine-led discovery about how a certain protein is activated in tumor cells. The finding, spearheaded by researchers with the School of Biological Sciences, could eventually lead to possible therapies for the especially dangerous melanoma and pancreatic adenocarcinoma, as well as the most common type of childhood brain cancer and adult skin cancer. A paper describing the project appears in Life Science Alliance.
The discovery concerns the GLI1 protein, which is important in cell development but has also been found turned on in various cancers. GLI1 is typically activated by the Hedgehog signaling pathway, known as HH. However, scientists have known for about a decade that crosstalk (interaction) between HH and the mitogen-activated protein kinase pathway has a role in cancers.
"In some cases, proteins in one pathway can turn on proteins in another," said lead author A. Jane Bardwell, project scientist in UCI's Department of Developmental and Cell Biology. "It's a complex system. We wanted to understand the molecular mechanism that leads to GLI1 being activated by proteins in the MAPK pathway."
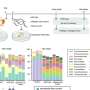
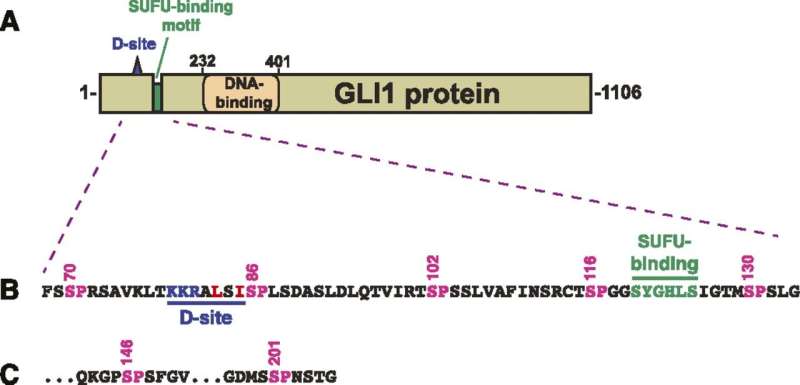
GLI1 normally binds tightly with a protein called SUFU. That protein suppresses GLI1, preventing it from penetrating cell nuclei and turning on genes. The scientists examined seven sites on the GLI1 protein that could be phosphorylated, or have a phosphate group transferred onto it.
"We identified three that can be phosphorylated and are involved in weakening the binding between GLI1 and SUFU," said Lee Bardwell, professor of developmental and cell biology, whose laboratory conducted the project. "This process activates GLI1, enabling it to enter the nucleus of cells, where it can cause uncontrolled growth resulting in cancer."
He noted that phosphorylation of all three sites causes a significantly higher level of GLI1 escape from SUFU than if just one or even two of them receive phosphate groups.
The discovery is a significant step toward more effective and personalized cancer treatments. "If we can understand exactly what is going on in a certain cancer or particular tumor, it could be possible to develop a drug specific to a specific tumor or individual patient," Bardwell said. "It would allow us to treat these diseases without the toxicity of basic chemotherapy." In addition, many tumors from the same cancer have different mutations among individuals. Eventually, it may be feasible to screen tumors to develop the best approach for each.
The research was performed in collaboration with the UCI School of Medicine's Department of Microbiology & Molecular Genetics and the Stanford University School of Medicine's Department of Dermatology.
More information: A. Jane Bardwell et al, ERK2 MAP kinase regulates SUFU binding by multisite phosphorylation of GLI1, Life Science Alliance (2022). DOI: 10.26508/lsa.202101353
Provided by University of California, Irvine