Scientists develop novel circulating tumor DNA biosensor
Nucleic acids analysis is mainly used in pathogen detection, genetic disease identification and early cancer diagnosis. For example, quantitative analysis of circulating tumor DNA (ctDNA), a free DNA fragment derived from malignant cells which carries tumor specific sequence changes, can help obtain abundant information about tumors, including gene point mutation, genome integrity. Therefore, ctDNA is considered as a personalized tumor marker and plays a key role in cancer diagnosis and malignancy evaluation.
Miao Peng's group from the Suzhou Institute of Biomedical Engineering and Technology (SIBET) of the Chinese Academy of Sciences has recently developed an electrochemical DNA nanomachine based on looped bipedal DNA walking reaction for highly sensitive analysis of nucleic acids. Relevant results were published in ACS Central Science.
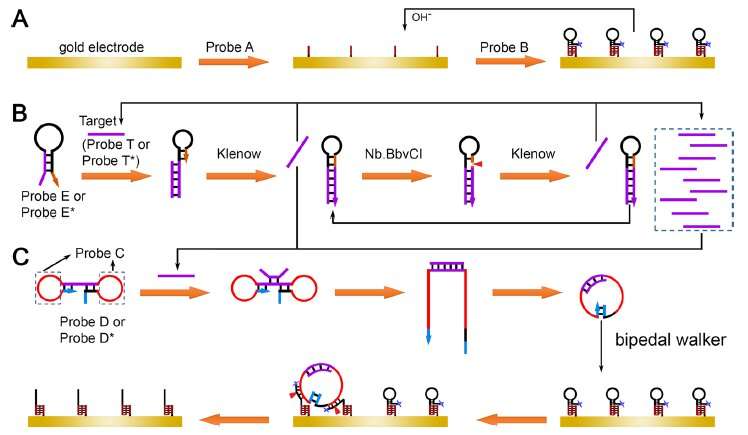
Miao and his team built a pH controllable intermolecular triple helix DNA nanostructure between DNA probes A and B through sequence design, and then constructed a renewable modified electrode interface.
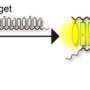
They designed a simple but effective strand displacement amplification strategy to amplify the information of a target sequence. "Through the integration of primer and template into a single hairpin structured DNA probe, the reaction rate was effectively improved," said Miao.
In the presence of a target sequence, a large number of single-stranded DNA products could be produced.
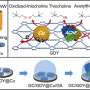
Furthermore, they developed a novel looped bipedal DNA walking strategy. The two loops of dumbbell structured DNA probe contained DNAzyme sequences, which could not react with the track strands at the electrode interface initially (inactivated state).
When it was activated by the single-stranded DNA produced by the above strand displacement amplification, a looped structured DNA probe was formed from dumbbell probe. The bipedal walkers were activated for further interaction with the track strands at the electrode surface, inducing the changes of electrochemical response.
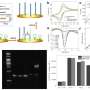
Based on the developed method, 2.2 aM (about 1.3 copy/μL) sensitivity could be realized under optimized experimental conditions, according to their results. "This method shows good selectivity," said Miao.
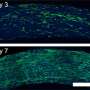
The clinical serum samples and throat swab samples were further tested and verified.
Through the analysis of abnormal electrochemical signals, the corresponding patients can be effectively identified from the healthy control groups, according to Miao.
The strategy also provides a fast and sensitive new way for detecting DNA markers of acute infectious diseases.
More information: Peng Miao et al, DNA Hairpins and Dumbbell-Wheel Transitions Amplified Walking Nanomachine for Ultrasensitive Nucleic Acid Detection, ACS Nano (2022). DOI: 10.1021/acsnano.1c11582
Journal information: ACS Central Science , ACS Nano
Provided by Chinese Academy of Sciences