RNA map of the mammalian cell nucleus reveals new insights into gene regulation
Most people are familiar with the cell nucleus from grade school biology as a storage compartment for DNA. But the nucleus also contains several distinct structures, called nuclear bodies or domains, whose roles scientists are just beginning to understand.
Some of these structures are brimming with genes' messages, also known as RNA transcripts.
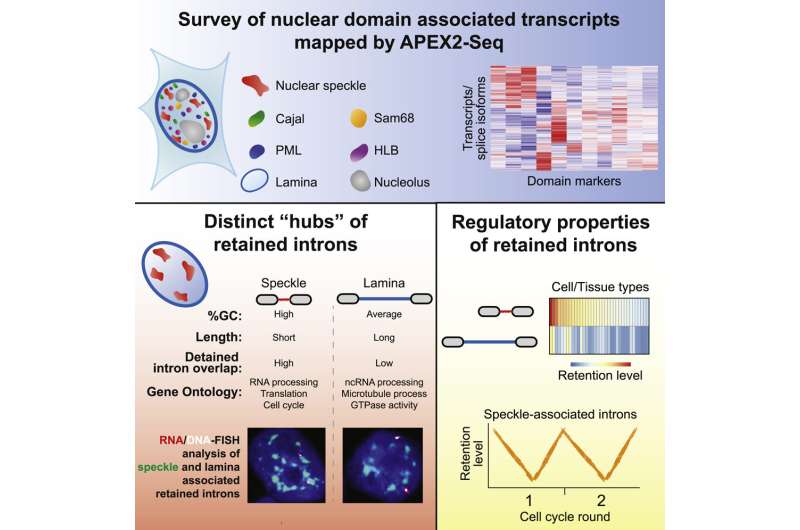
Now, U of T researchers have reported the first large-scale survey of RNA transcripts that are associated with different nuclear bodies in human cells. Their work suggests that these structures act as hubs to coordinate gene regulation and cell division.
"It was known that some nuclear domains contain RNA, but the composition of that RNA was not systematically probed in previous studies," said Benjamin Blencowe, senior author on the study and a professor of molecular genetics in the Donnelly Centre for Cellular and Biomolecular Research, at the Temerity Faculty of Medicine.
"Our data has shed light not only on the RNA composition of different nuclear domains, but also provides clues as to the functions of some of these domains," he said.
Molecular Cell published the findings.
Until now, the information on nuclear body composition has trickled in piecemeal because there were no methods enabling a systematic survey of RNA localized to these structures. But postdoctoral fellow Rasim Barutcu and graduate student Mingkun Wu realized they could apply a method called APEX-Seq, which had been developed by Stanford and Berkeley scientists.
APEX is an enzyme that can be fused to any protein of interest and allows labeling of RNAs, and other biomolecules, in its proximity. The labeled RNAs can then be isolated and identified by sequencing. By fusing APEX to various marker proteins residing in the different nuclear bodies, Barutcu and Wu were able to create RNA maps for each. In this effort, they collaborated with Ulrich Braunschweig, a senior research associate in the Blencowe lab, and with the groups of Anne-Claude Gingras, a U of T professor of molecular genetics and a senior scientist at the Lunenfeld-Tanenbaum Research Institute, at Sinai Health System, Philipp Maass , a U of T assistant professor of molecular genetics and scientist at The Hospital for Sick Children, and Robert Weatheritt, a principal investigator at the Garvan Institute of Medical Research, Australia.
The team discovered swaths of novel RNAs, from several hundred to thousands, across the nuclear bodies. Previously, only a handful of transcripts were known to be associated with some of these structures, said Barutcu, whose research was supported by the Banting Postdoctoral Fellowship and a fellowship from the Canadian Institutes of Health Research (CIHR).
One piece of data immediately struck the researchers. The nuclear bodies known as the speckles were associated with surprisingly high numbers of RNA transcripts with retained introns, segments which do not code for proteins.
When a gene is transcribed into RNA, introns must be spliced out in the nucleus before the transcript can be released into the cell's interior to serve as a template for making proteins.
The finding led them to realize that speckles are associated with a class of introns with delayed splicing. The nature of the transcripts provided a clue to their function. They were transcribed from genes that control various aspects of gene regulation and the cell division cycle. Genes controlling cell cycle progression must be activated in a timely manner so that their protein products are made only when they are needed. Errors in this process are well known drivers of cancer.
The researchers came up with a model in which the role of the speckles might be to coordinate intron removal from transcripts in order to regulate their release from the nucleus, and their subsequent translation into protein factors required for gene regulation and the cell cycle. This mechanism would help ensure a rapid response to cellular signals to make the right kinds of proteins at the right time.
Furthermore, when speckles were disrupted, this altered the splicing of the retained introns, including those located in genes that are directly involved in control of the cell cycle, supporting the idea that the speckles are linked to cell cycle progression.
The model opens up new ways of thinking about cell cycle regulation with implications for cancer research, said Blencowe, who holds Canada Research Chair in RNA Biology and Genomics and Banbury Chair in Medical Research.
"We've uncovered a mechanism involving differential intron retention linked to speckle integrity that could play an important role in not just normal cell division but also how it goes wrong in cancers," he said.
In addition to the speckles, the team also found large numbers of intron-retained transcripts associated with the nuclear lamina, which forms at the periphery of the nucleus, but the functional significance of this observation remains unclear.
The researchers said they hope others in the field will take advantage of their datasets and open new avenues of research into nuclear body function where many questions remain open.
Blencowe also added that the project was made possible by the freedom to pursue new directions afforded by the now scrapped CIHR Foundation grant scheme that provided long-term research funding.
More information: A. Rasim Barutcu et al, Systematic mapping of nuclear domain-associated transcripts reveals speckles and lamina as hubs of functionally distinct retained introns, Molecular Cell (2022). DOI: 10.1016/j.molcel.2021.12.010
Journal information: Molecular Cell
Provided by University of Toronto