Bacteria prompt a new look at the dynamics of collective behavior

A study led by Princeton researchers has revealed how bacteria navigate obstacles to ensure cohesive group movement. The finding has implications for understanding the general processes of collective migration, from cancer cells responding to chemical stimulus to wildebeest moving across the savanna.
Principal investigator Sujit Datta said he had noticed that migrating bacteria populations tended to stay in well-organized formations even when they encountered obstacles. To tease apart what was happening, his team created conditions that might cause the groups of bacteria to splinter into winners and losers.
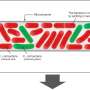
Datta likened his team's experimental setup to a foot race with a squiggly starting line, where some individuals start in front of others, giving them an advantage. With a rich vein of nutrients ahead, the bacteria cells were motivated to speed forward and gobble up as much food as possible.
"To our surprise, what we found was that the cells that started out ahead moved slower, and the cells that started out behind moved faster, so the cells behind were always able to catch up," said Datta, an assistant professor of chemical and biological engineering. "Eventually those squiggles we had initially forced into the shape of the population smoothed out as the cells continued to move together."
This study, published March 8 in the journal eLife, shows how individual self-regulation leads to a profound, emergent collective phenomenon that cuts against common assumptions about individual survival strategies.
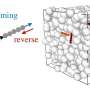
It follows earlier work from Datta's lab that established how bacteria spread like a wave traveling outward as they seek and consume nutrients in their surroundings, moving together as a group despite encountering the many obstacles in the wave's path. But whereas the earlier study focused on the basic effect of obstacles on group movement, the more recent work focuses on how populations stay together even when they encounter significant disruptions to the group as a whole.
Datta's race among a population of Escherichia coli took place in a hydrogel matrix, like an obstacle course for bacteria. The researchers 3D-printed the cells in defined arrangements and then used a confocal microscope to track the movement of the cells, genetically engineered to glow, as they ate their way through the matrix.
"By using this microscope, you have a very precise, pin-point accurate measurement of light in three dimensions," said Daniel Amchin, a graduate student in chemical and biological engineering and one of the paper's authors. With a long sequence of high-tech snapshots, the team developed an elaborate view of how the cells move—individually and as a group—over time.
They initially considered that the smoothing they observed was the result of the cells simply moving randomly. "But when you do the math," Amchin said, "the random component of their motion couldn't possibly cause the squiggles to smooth out as fast as we observed them to."
That's when the team developed an alternative explanation, supported by their experiments, simulations and mathematical theory.
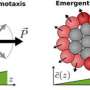
The leading cells had access to the most food because they were on the front edge, closest to the next yummy bite. Followers had correspondingly less, depending on their starting position. To the researchers' surprise, an individual cell's response to the nutrient gradient became less effective the more it ate. The leading cells, increasingly saturated, slowed down. The followers kept apace, meaning they got more food than their starting positions would have indicated without this emergent distributive phenomenon. The effect was that the migrating population stuck together despite the obstacles and incentives that might have split them apart.
"That's the beauty of this," Datta said. "The behavior of these individuals is tuned, perhaps through million years of evolution, to favor collective outcomes even though the individuals have no concept of the other cells." An analysis from Datta's group, forthcoming in Physical Review Letters, further supports this idea, establishing quantitative predictions for the conditions under which populations should be able to stick together—and showing that naturally occurring populations tend to satisfy these conditions.
These insights lend a detailed and rigorous framework to the general understanding of collective migration, beyond bacteria, such as how herding, flocking and schooling organisms—and even microscopic swimming robots—coordinate their actions in large groups.
"We think this is a more generic principle describing how populations of many different moving entities can manage to travel together as a group, since the ability of other cells or agents to sense and respond to various stimuli is inherently limited in many cases," Datta said. "So, we hope that our work will motivate other people to study this effect in other contexts."
More information: Tapomoy Bhattacharjee et al, Chemotactic smoothing of collective migration, eLife (2022). DOI: 10.7554/eLife.71226
Ricard Alert et al, Cellular Sensing Governs the Stability of Chemotactic Fronts, arXiv:2107.11702v1 [physics.bio-ph]
doi.org/10.48550/arXiv.2107.11702
Journal information: eLife , Physical Review Letters
Provided by Princeton University