New genome analysis method and model address blind spot towards uncommon species in mixed samples
For higher accuracy and more effective microbial diagnostics, researchers at the Earlham Institute have evaluated a new approach to analyze metagenomic samples that may contain low numbers of certain species—helping to prevent crucial genomic information from being overlooked or lost.
Existing methods have created a 'biodiversity blindspot' towards the rare; meaning low abundance species may be underrepresented, while genetic variants could be missed altogether. This current oversight could, for example, fail to spot crucial information about resistance to antibiotics or delay the point at which the emergence of a new strain becomes detectable.
By combining advanced sampling techniques with mathematical models, using Oxford Nanopore Technology's software, Earlham Institute scientists were able to amplify and enrich metagenomic samples to reveal crucial details that may have been unidentified before.
Metagenomics approaches are used to analyze genomic function and diversity of complex microbial communities from environmental, agricultural and clinical settings.
The whole-genome shotgun approach in metagenomics enables the reliable identification of organisms down to species and strain level. This allows microbiologists to evaluate bacterial diversity and detect the abundance of microbes in various environments.
A popular tool for understanding mixed species, metagenomics is particularly used for analyzing individual species (that cannot be isolated from their environment), or gene clusters such as antibiotic resistance genes—to shed light on genomic function or generate reference sequences for unculturable organisms.
With the increasing use of long read technologies, either on their own, or combined with short-read technologies, metagenome assembled genome (MAG) contiguity and accuracy metrics have improved. Such approaches have been widely applied including assembling pathogen genomes from clinical samples, bacterial genomes and gene clusters from the human gut to the rumen microbiome of cattle.
Yet, despite these successes, difficulties still remain about the reliability of MAG approaches when faced with complex populations. As a solution, scientists at the Earlham Institute applied a new approach called adaptive sampling, which is a method for software-controlled enrichment unique to nanopore sequencing platforms.
By creating a model and evaluating the use of adaptive sampling approaches, the team tested its potential for enrichment of rarer species within metagenomic samples—producing a synthetic mock community and constructed sequencing libraries with a range of DNA molecule lengths. The team also tested model communities of gut microbes and evaluated performance on complex microbial community samples taken from a garden compost heap.
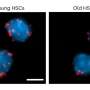
Study lead and Group Leader at the Earlham Institute Dr. Richard Leggett, says that "metagenomic samples are composed of a range of different species at varying levels of abundance. This can mean that sequencing these samples produces data resulting in deep coverage of some species; with low or partial coverage of others."
"For rarer species, this is likely to result in much poorer assemblies and a reduction in the ability to distinguish between strains or related species. Effective enrichment strategies to maximize the sequence outputs of the rare species would address this weakness, and biodiversity blindspot."
Adaptive sampling offers a potential solution to enrich species of interest in metagenomic samples. Using a relatively simple library construction method, samples can be prepared for sequencing within an hour without the need for amplification. However, a challenge for microbiome research is the difficulty of extracting long intact DNA molecules from complex metagenomic samples.
"Reducing host DNA is an important consideration in many diagnostic applications," he says, "especially in clinical settings. We wanted to investigate the effect of DNA molecule length on the efficiency and efficacy of adaptive sampling to determine its usefulness for both MAG and diagnostic applications."
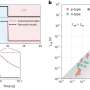
First author and Postdoctoral Scientist at the Earlham Institute Dr. Samuel Martin, says that "in our study, we present a mathematical model that can predict the enrichment levels possible in a metagenomic community given a known relative abundance and read length distribution. Through Oxford Nanopore Technology's adaptive sampling approach, we demonstrated enrichment in terms of both yield and composition, in a synthetic mock metagenomic community and in a complex real sample."
"We found that enrichment was higher for less abundant species, and for libraries with a higher average molecule length, showing that extraction methods that can preserve molecule length are key to obtaining the highest enrichment."
"By performing targeted enrichment on a low abundance species, we were able to significantly reduce the time taken to achieve a high accuracy, single-assembly, compared to non-targeted sequencing. We expect that adaptive sampling will prove to be a useful tool for many nanopore-based metagenomic studies."
The research is published in Genome Biology.
More information: Samuel Martin et al, Nanopore adaptive sampling: a tool for enrichment of low abundance species in metagenomic samples, Genome Biology (2022). DOI: 10.1186/s13059-021-02582-x
Journal information: Genome Biology
Provided by Earlham Institute