Protein domain structures affect the quality of stem cells
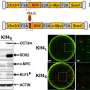
The original iPS cells were reprogrammed by transducing four genes: OCT3/4, SOX2, KLF4, and c-MYC. Later experiments would show that c-MYC can be replaced with MYCL. Both are MYC proteins and thus oncogenes, but animal experiments have demonstrated that iPS cells reprogrammed using MYCL are less likely to cause tumors than iPS cells reprogrammed with c-MYC. Furthermore, MYCL enhances the reprogramming efficiency, but why is unknown. A new study from CiRA reports that two domains in the MYC proteins, MYC Box domain 0 (MB0) and MB2, are responsible.
Reprogramming factors initiate a series of events inside a cell that leads to the formation of iPS cells. To understand why c-MYC and MYCL result in different reprogramming efficiencies, Chiaki Akifuji, a Ph.D. candidate who is the first author on the study, presumed specific units in the amino acid sequences of the two proteins were the cause.
"MYC proteins have six MYC Box domains. These domains are responsible for protein binding. We hypothesized that the domains behave differently in c-MYC and MYCL to cause the different reprogramming efficiencies," she said.
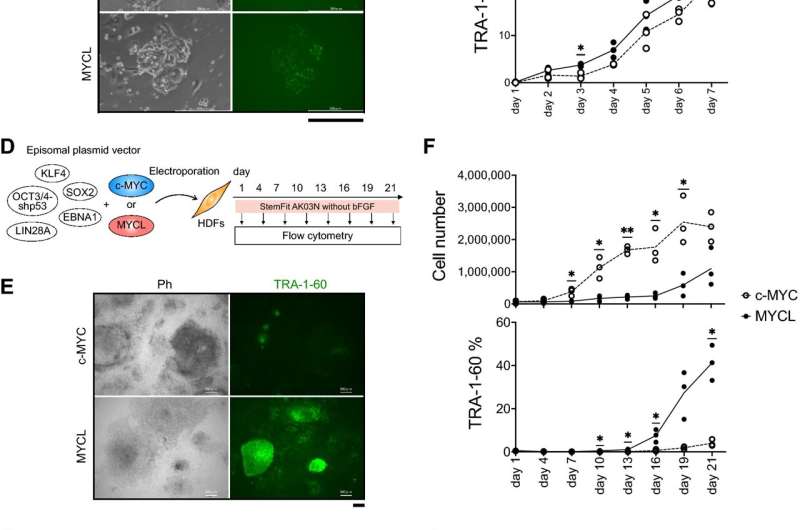
The study reprogrammed human fibroblasts using c-MYC and MYCL mutants in which each MB domain was deleted. While all deletions in MYCL resulted in fewer iPS cells, deleting either MB0 or MB2 prevented any reprogramming from happening at all. This was not the case for deleting MB0 in c-MYC, suggesting that MB0 has a different function in the two MYC proteins.
"The different functions are surprising because the domains are well conserved," said Akifuji.
Further experiments discovered that MYCL MB0 promotes the expression of proteins responsible for cell adhesion, suggesting MYCL affects how reprogramming cells bind to other cells.
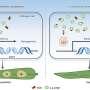
During reprogramming, cells undergo a mesenchymal-to-epithelial transition. This transition also occurs during natural development and cancers. Cell behavior, such as that during reprogramming, depends on how the cell interacts with its environment, which is affected by the mesenchymal-to-epithelial transition.
On the other hand, MYCL MB2 was found to bind to several RNA-binding proteins that regulate how RNA is distributed and translated in a cell. This effect determines where a protein is located in the cell and whether it is activated, which also potentially influences reprogramming.
Thus, the study suggests that the efficient reprogramming by MYCL depends on two domains that work synergistically by regulating cell adhesion and RNA processing to promote the mesenchymal-to-epithelial transition.
"Our goal is to fully elucidate the reprogramming mechanism. Understanding each factor that contributes will help us generate iPS cells of clinical grade at lower cost," said CiRA Junior Associate Professor Masato Nakagawa, who managed the study.
More information: Chiaki Akifuji et al, MYCL promotes iPSC-like colony formation via MYC Box 0 and 2 domains, Scientific Reports (2021). DOI: 10.1038/s41598-021-03260-5
Journal information: Scientific Reports
Provided by Kyoto University