Researchers discover new hiding place for antibiotic resistance
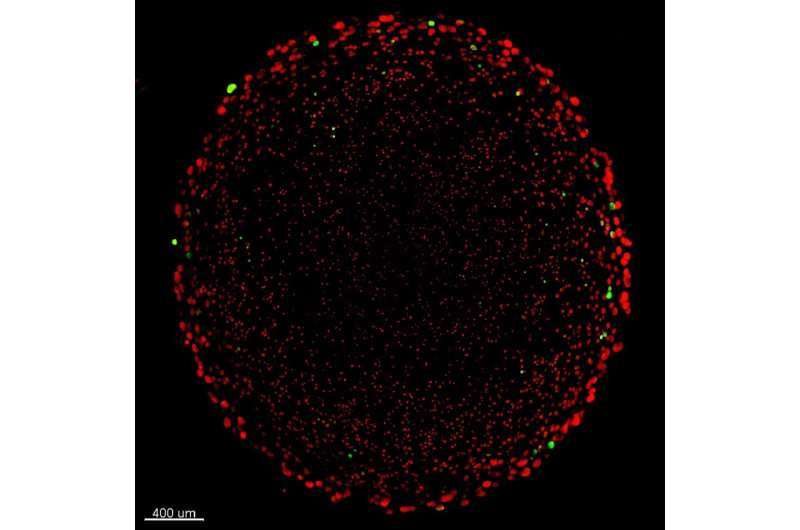
Genes that make bacteria resistant to antibiotics can persist longer than it was previously believed. This was recently shown in a new University of Copenhagen study that reports a previously unknown hiding place for these genes. The finding represents a new and important piece in the puzzle to understand how bacterial antibiotic resistance works.
Antibiotic resistance is a race between us humans, who strive to find new antibiotics that can treat infectious diseases—and bacteria, which are becoming increasingly resistant. For now, bacteria are way ahead, which is why it is important for us to learn more about antibiotic resistance. A Danish research group has discovered a new piece of the puzzle that helps us better understand the 'enemy."
University of Copenhagen researchers have shown that the prevailing assumption that resistant bacteria lose their resistance capability when antibiotics are not present is a truth requiring significant modifications.
"One widespread strategy to combat antibiotic resistance has been to use antibiotics for a period of time and then take a break. The belief is that resistant bacteria will lose their resistance genes or be outcompeted during the break, after which the antibiotics will work again. But that approach doesn't seem to hold up," says co-author Associate Professor Mette Burmølle of the Department of Biology.
Co-first author Henriette Lyng Røder elaborates: "Our study demonstrates that resistance genes are able to hide in inactive bacteria, where they form a hidden reserve of resistance that bacteria can rely on. In other words, they don't just disappear when antibiotics aren't around."
Biofilm deals resistance genes a strong card
Most bacteria live and interact in what are known as biofilms—where microbial communities are encased in a matrix of mucus they form, often on the surface of a material. Biofilms are found everywhere from stones and plants, to plaque on the teeth, to implants. Biofilms contain both active and inactive bacteria. The mucus and hibernation of inactive bacteria make biofilms a fortress able to withstand large amounts of antibiotics. But the new study shows that biofilms deal bacteria another strong card.
"We can see that the active bacteria living nearest the outer edge of the biofilm lose resistance genes when antibiotics aren't present. However, deeper within the biofilm, there is a layer of inactive bacteria which hibernate away safely. These carry resistance genes even if they don't need them. This is important because it means that biofilms can essentially act as a reserve for the storage of many types of resistance genes," explains Urvish Trivedi, the study's co-first author.
Resistance genes are typically spread by small DNA molecules that transfer between the bacteria they use as hosts. Until now, it was thought that bacteria only keep plasmids for as long as they can benefit from them, e.g., by the resistance genes plasmids carry, or else lose them. Indeed, plasmids are no free lunch. They steal energy from a bacterium and cause it to grow more slowly. And since active bacteria are in constant competition with each other, it has been a mystery as to why many bacteria carry around plasmids without doing them much good—known as selection.
The new study provides one of the answers. When it comes to inactive bacteria, conditions are different.
"In contrast to the active bacteria in biofilm, inactive bacteria in biofilm don't grow. As such, they don't compete. This allows space for them to carry plasmids. In this way, a reserve of resistance genes is built up into biofilm. Obviously, it's a huge advantage for bacteria to be able to save resistance up to 'bad times' – in this case, when a bacterium meets an antibiotic," explains Mette Burmølle.
We're not getting rid of them
The researchers estimate that resistance stocks in biofilms are primarily built up in environmental bacteria, found in soil, air and wastewater among other places. However, it is well established that different species of bacteria can transmit resistance to each other. For example, resistance in environmental bacteria can be transmitted to the types of bacteria that make people sick.
"An enormous number of bacteria with antibiotic-resistant genes derived from humans and livestock end up in sewage and may spread along that path into the environment. One concern is that such bacteria could end up turning environmental bacteria into pathogens—bacteria that cause disease. In this way, everything is connected," says Jonas Stenløkke Madsen, another senior author of the study.
All in all, the new findings inform us that resistant bacteria are even better at surviving than we thought. Madsen concludes:
"In the bigger picture, this means that if there are a lot of inactive bacteria in the environment, in soil for example, then resistant genes don't just gradually disappear when antibiotics aren't present. Therefore, we ought to consider abandoning the idea that we can get rid of resistance genes and instead assume that they are always present. Understanding these dynamics can better equip us to battle antibiotic-resistant bacteria."
The research results are published in the scientific journal npj Biofilms and Microbiome.
More information: Henriette Lyng Røder et al, Biofilms can act as plasmid reserves in the absence of plasmid specific selection, npj Biofilms and Microbiomes (2021). DOI: 10.1038/s41522-021-00249-w
Provided by University of Copenhagen