New point spread functions developed for particle 3D tracking
Nanoparticle positioning and tracking have a wide range of uses in life science, drug research and development. Real-time recording of the intracellular and extracellular motion of nano particles is of great significance in exploring the basic laws of life activities and drug transformation as it is crucial for clarifying key scientific issues such as disease pathogenesis, viral dynamic infection of host cells, and promoting the development and transformation of nano-drugs.
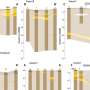
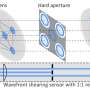
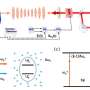
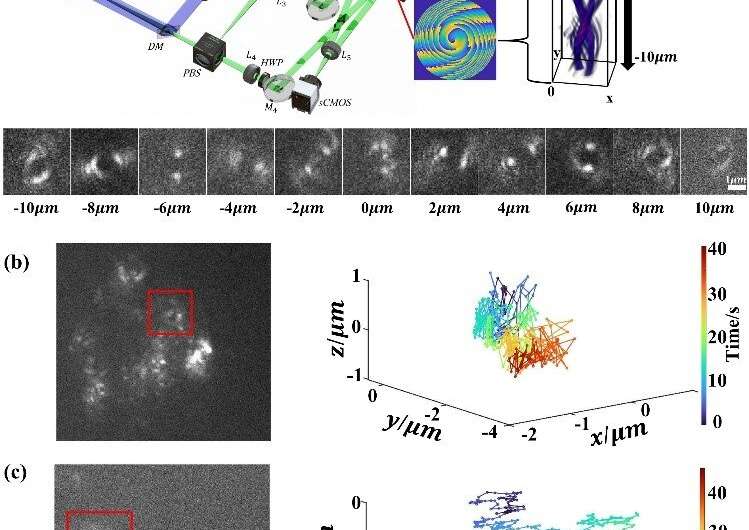
In a new research published on Optics Letters, researchers led by Prof. Zhang Yunhai from the Suzhou Institute of Biomedical Engineering and Technology (SIBET) of the Chinese Academy of Sciences (CAS) modulated the phase of the emitted light to reshape the point spread function (PSF) and thus encode the axial position of the emitter. The information on the axial position of the particles can be obtained by estimating the variation of the shape or size of the PSF.
The researchers designed two new point spread functions: 2π-double helix point spread function (2π-DH-PSF) and splicing exponential function point spread function (SE-PSF), and used them to three-dimensional (3D) track biological particles.
The 2π-double-helix PSF (2π-DH-PSF) which can rotate 2π radians can track particles three-dimensionally in the axial range of 10 μm. Combined with the defocus phases, and adopt a specific optical setting, which results in a final DH-PSF rotation angle of 720 degrees, can achieve a fourfold extension of the depth of field compared to a conventional DH-PSF.
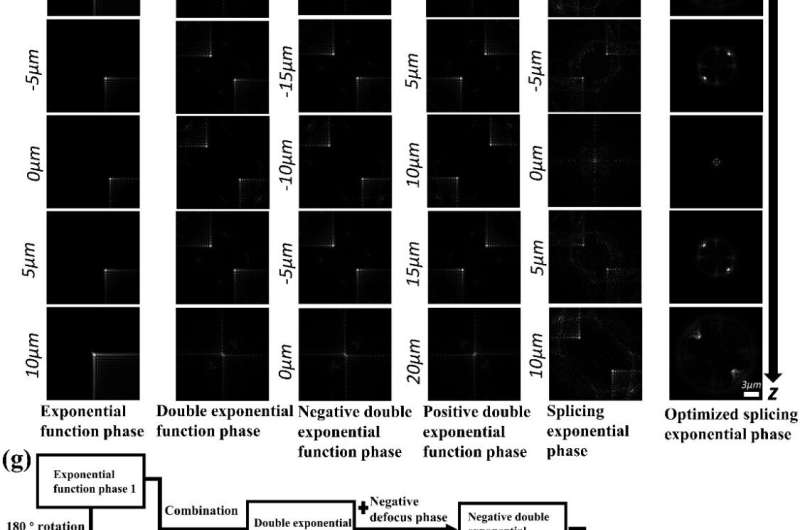
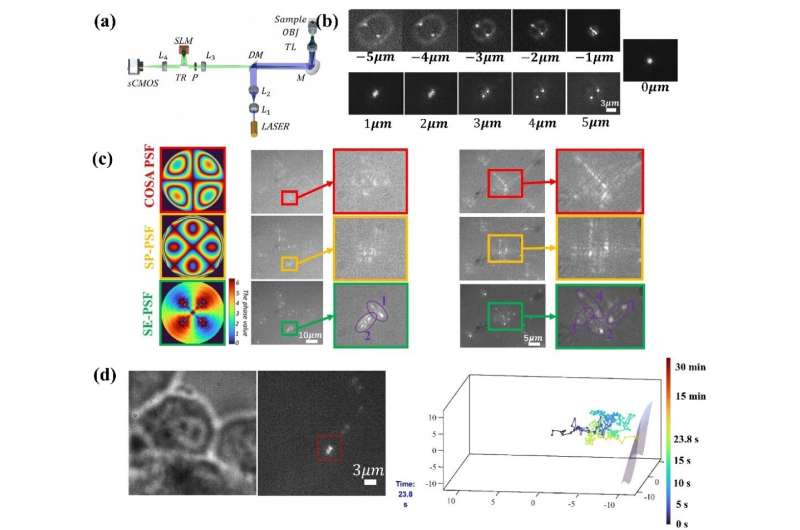
The SE-PSF can control the spatial extent and axial detectable range by adjusting the design parameters. Taking exponential function phase and defocus phase as basic units, the optimized phase of SE-PSF generated by splicing, symmetry, optimization and other operations can track particles in three dimensions within 20μm axial range.
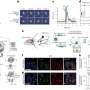
The SE-PSF with a smaller spatial extent can effectively reduce the overlap of nanoparticle images and realize the 3D localization of dense multi-particles.
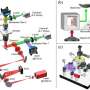
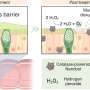
The 3D tracking technology for particles can record the trajectory of the virus in the extracellular biological gel (such as mucus), and the process of virus particles entering living cells.
"It can be used to calculate particle's average velocity, diffusion coefficient, etc. Therefore, it can provide a reference for studying the dynamic transport process of virus particles infecting host cells," said Prof. Zhang Yunhai, the team leader.
In addition to the three applications of outer membrane vesicles, viruses, and nano-drug carriers, it can also be applied to neural vesicles (50–500 nm), chylomicrons (75–600 nm), and chromosomes (30–750 nm).
The tracking and positioning technology provides new research ideas and methods for dynamic process of neurotransmitter signal transduction, digestion and absorption of nutrients in the gastrointestinal tract, and the replication of genetic material, according to Prof. Zhang.
More information: Hangfeng Li et al, Particles 3D tracking with large axial depth by using the 2π-DH-PSF, Optics Letters (2021). DOI: 10.1364/OL.434981
Journal information: Optics Letters
Provided by Chinese Academy of Sciences