Artificial intelligence accelerates search for markers of resistance to sugarcane yellow leaf disease

Yellow leaf disease, a major sugarcane pest in Brazil, is caused by a virus resistant to thermal treatment. An infected plantation can be saved only by growing plantlets in tissue culture in the laboratory and planting them out, a time-consuming process that requires specialized infrastructure and personnel. According to a group of scientists who have long studied the problem, the most effective way to control the disease is to develop varieties that are resistant to the sugarcane yellow leaf virus. This is the purpose of a project that is being conducted with FAPESP's support.
The scientists are affiliated with universities (University of Campinas—UNICAMP, São Paulo State University—UNESP), and state-run agribusiness research institutes (Campinas Agronomic Institute—IAC, Biological Institute—IB) from São Paulo state, Brazil, and with Ecuador's ESPOL Polytechnic. In an article published recently in the journal Scientific Reports, they describe how they used genomics, machine learning, and statistics to refine and accelerate the search for molecular markers of resistance to the disease.
The group found that resistant varieties are mostly types of so-called energy cane, with higher fiber content and less sugar content than conventional cane, and hence fittest for second-generation or cellulosic ethanol production, although at least one is fit for sugar or conventional ethanol production—so much so that it has just been commercially launched by IAC.
The researchers evaluated 97 sugarcane genotypes, including wild germplasms of Saccharum officinarum, S. spontaneum and S. robustum, traditional sugar and energy cane clones, and commercial varieties developed by Brazilian breeding programs.
"We analyzed the resistance of each of these varieties to yellow leaf disease. The aim was to associate disease resistance with genetic traits. We used several different molecular markers, which are DNA variations, deploying next-generation sequencing to access this information," said Ricardo Pimenta, a doctoral researcher at UNICAMP's Center for Molecular Biology and Genetic Engineering (CBMEG).
The varieties were selected by IAC's cane breeding program. Most were part of the program, but there were also specimens from the Inter-University Network for Development of the Sugar and Ethanol Industry (RIDESA) and the Sugarcane Technology Center (CTC).
"The collection was representative of the variability of Brazilian cane, both planted and used in cross-breeding to produce other varieties," said Anete Pereira de Souza, a professor in the Department of Plant Biology at UNICAMP's Institute of Biology and project coordinator at CBMEG.
The symptoms of sugarcane yellow leaf disease typically appear in the later stages of the plant's development. The main symptom is intense yellowing of leaf midribs. The disease alters photosynthetic efficiency as well as sucrose metabolism and transport, impairing growth and yield.
Methods
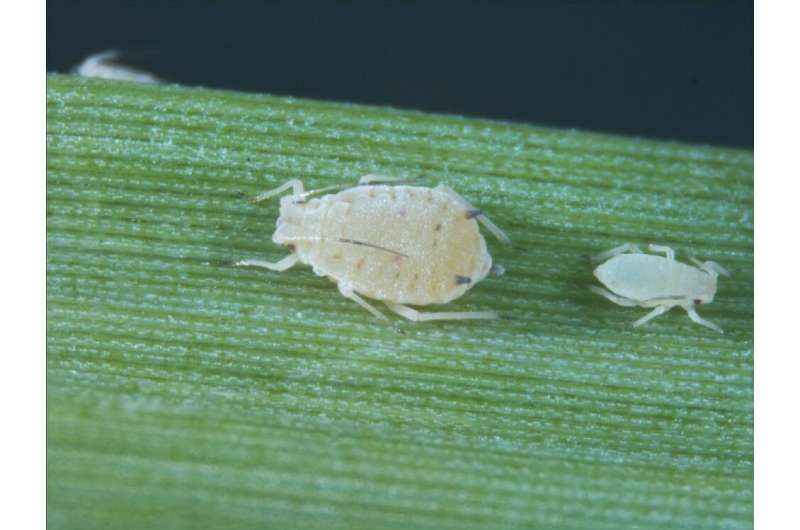
According to Pimenta, the article describes an experimental procedure that had never been attempted before. "The yellow leaf virus is transmitted by the sugarcane aphid Melanaphis sacchari," he said. "The IAC team planted experimental plots and at the same time reared aphids on plants already infected by the virus. The aphids were then released into the uninfected plantations and monitored in a controlled process of inoculation and infection. Previous studies used a similar approach, except that they planted the cane and left it to be infected naturally, as it were, in a less controlled inoculation process."
Viral load was measured by reverse transcription-quantitative polymerase chain reaction (RT-qPCR). "We used PCR to quantify the virus in the entire set of cane varieties analyzed," Pimenta explained. Disease severity was assessed by observing leaf yellowing intensity and other symptoms, which can often be elusive.
To establish associations between disease resistance and genetic traits, the researchers used genomic techniques, machine learning (an artificial intelligence procedure based on pattern recognition), feature selection, and marker-trait association methods.
"What we normally find in genomic association studies is markers that strongly influence the phenotype [observable characteristics]. That's a problem because you can't find the others that have less influence on the phenotype. Feature selection captures markers that influence the phenotype in a narrower sense, so we used the technique to screen molecular markers more efficiently," said Alexandre Hild Aono, also a researcher at CBMEG.
Machine learning was used to construct a model that predicted whether a variety was resistant or susceptible to the virus, given the specified genetic markers. "To do this, the algorithms first have to rate markers 'highly important' or 'less significant' for predictive purposes. The question therefore was this: If the system rates certain markers highly, do they also influence the phenotype? Our investigation confirmed this was indeed the case," Aono said. "We combined the various methodologies and succeeded in filtering and selecting markers with the most potential to influence configuration of the disease directly, even though they may not be especially conspicuous [in terms of influencing the phenotype]."
One of the aims of the study, Souza recalled, was to compare the results of the methodological techniques in order to see if they converged. "We found a larger set of markers with the methodology proposed, but it was also validated by traditional statistics. They talk to each other, and by using them in association we were able to obtain a much broader dataset for analysis and a richer basis for genetic improvement," she said.
The multidisciplinary team achieved an unprecedented level of detail in this study, she added: "We compared the methodologies and demonstrated the efficiency and necessity of using a more refined statistical approach. Nothing in the literature suggests anyone had ever done this type of analysis, or reared the aphids, infected the plants, and then measured viral load with quantitative PCR. The results serve as a foundation and guide for future research, contributing to a better understanding of the molecular mechanism involved in the disease."
The study was supported by FAPESP via a master's scholarship and a direct doctorate scholarship awarded to Pimenta; a direct doctorate scholarship awarded to Aono; and a postdoctorate scholarship awarded to Carla Cristina da Silva, another co-author of the article.
Disease resistance
According to Pimenta, genuinely resistant plants that do not manifest symptoms or accumulate viral load account for a small proportion of the total. "Few are really resistant. Most plants don't display symptoms but accumulate viral load, which eventually becomes a problem because the pathogen is there without the producer being aware of it. Our findings can help eliminate susceptible varieties as well as tolerant varieties, which accumulate viral load without displaying symptoms and can become a viral reservoir," he said, stressing that the main challenge in crop breeding is selecting the best varieties without losing too much variability.
Many resistant varieties are representatives of energy cane. "They have a recent 'wild' ancestor that's more disease resistant, so this result isn't a surprise. But we also discovered more commercial varieties that proved resistant, and these are more interesting. One of them, IACSP04-6007, proved so promising that it's just been launched by IAC's breeding program," Pimenta said.
Besides this one, the following varieties were also found to be most resistant to yellow leaf disease (no symptoms and low viral loads): IACBIO241, IACBIO257, IACBIO266, IACBIO270, IACBIO271, IACBIO273, IACBIO275, IACBIO279, IACCTC 05-3616, IACSP04-2510, IACSP98-5046, IJ76293, IN8482, IN8488, and Krakatau.
Pimenta also noted the importance of the genes associated with the markers identified. "Some of the most striking examples include the gene for a peroxidase, an enzyme previously associated with resistance to this disease; the gene for a Dicer, a very important enzyme in plants' viral response mechanism; and several genes containing leukin-rich repeats widely involved in plants' immune responses to pathogens," he said.
More information: Ricardo José Gonzaga Pimenta et al, Genome-wide approaches for the identification of markers and genes associated with sugarcane yellow leaf virus resistance, Scientific Reports (2021). DOI: 10.1038/s41598-021-95116-1
Journal information: Scientific Reports
Provided by FAPESP