Structural biology: Mechanisms of novel anti-cancer drugs elucidated
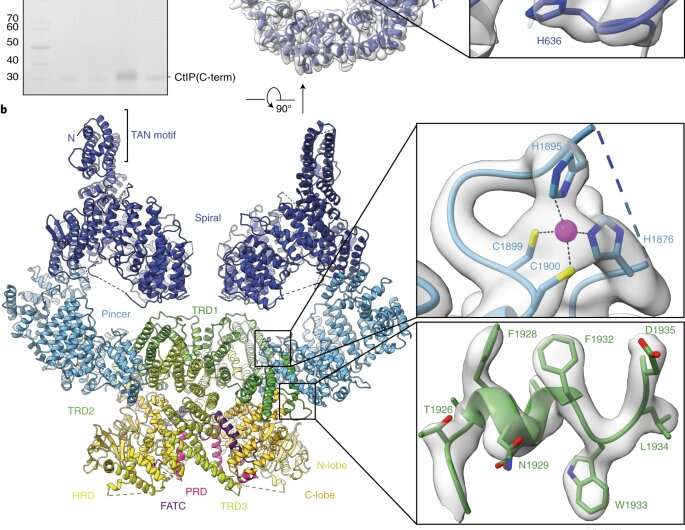
Double-stranded breaks (DSBs) are among the most hazardous forms of DNA damage. The checkpoint protein kinase ATM plays a key role in the repair of DSBs. Many anti-tumor drugs act by inducing DSB formation, so inhibition of ATM should enhance the sensitivity of cancer cells to these agents.
Owing to its instability and conformational flexibility, ATM—like other members of the family of checkpoint kinases—has been inaccessible to high-resolution structural analysis. However, detailed structural information greatly facilitates for the development of targeted and selective anti-cancer agents.
With the aid of cryo-electron microscopy, Karl-Peter Hopfner's research group, in collaboration with the drug company Merck, has now elucidated the structure of ATM at a resolution that enables a virtually complete atomic model of the protein to be built. Moreover, the model allows the binding modes of novel ATM inhibitors that are currently in clinical trials to be determined. The structural model can account for the selectivity of two ATM inhibitors, and thus provides the basis for the stereochemical optimization of new therapeutic compounds.
The study is published in Nature Structural & Molecular Biology.
More information: K. Stakyte et al, Molecular basis of human ATM kinase inhibition, Nature Structural & Molecular Biology (2021). DOI: 10.1038/s41594-021-00654-x
Journal information: Nature Structural & Molecular Biology
Provided by Ludwig Maximilian University of Munich