Profiling gene expression in plant embryos, one nucleus at a time
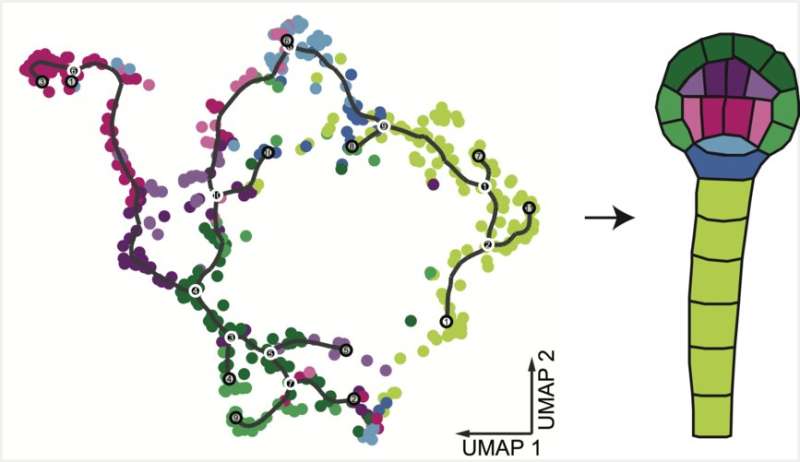
Following fertilization, early plant embryos arise through a rapid initial diversification of their component cell types. As a result, this series of coordinated cell divisions rapidly sculpts the embryo's body plan. The developmental phenomenon in question is orchestrated by a transcriptional activation of the plant genome. However, the underlying cellular differentiation programs have long remained obscured as the plant embryos were hard to isolate. In fact, previous attempts at creating datasets of the plant embryonic differentiation programs were incapable of overcoming two main obstacles: Either the information gathered lacked cell specificity, or the datasets were contaminated with material from surrounding non-embryonic tissues. Now, a team of Ph.D. students from GMI Group Leader Michael Nodine's lab has developed a method to profile gene expression at the single cell level in Arabidopsis embryos.
The authors, led by Ping Kao in collaboration with Michael Schon from the Nodine group, use an approach that consists of coupling fluorescence-activated nuclei sorting together with single-nucleus RNA sequencing (snRNA-seq). Hence, they sort the individual cells in an early plant embryo and sequence the messenger RNA within the individual nuclei. This provides insights into the various transcription profiles, or transcriptomes, within each cell in the plant embryo. With this approach, the team was capable of surmounting the obstacles that undermined previous attempts at creating gene expression atlases in plant embryos.
To explain their team's unique approach, Michael Schon from Nodine's group readily finds a striking parallel: "If you put a hamburger in a blender, it still has all the same components in all the same ratios, but you lose critical information about the burger's spatial organization. Elements essential to 'burger-ness' exist in the organization of the burger's parts, and these elements are lost in a 'burger smoothie.'" Schon elaborates, "Earlier methods consisted of grinding entire plant embryos into a 'plant embryo smoothie'; these were still useful in telling us the molecular components of the plant embryo and their ratios, but important information about the organism's organization was lost. Our transcriptome atlas is an effort to restore the information that most likely got 'averaged-out' in previous attempts."
Using this approach, the Nodine group was able to show gene expression patterns that could clearly distinguish the early Arabidopsis embryonic cell types. Consequently, the current work opens the door for uncovering the molecular basis of pattern formation in plant embryos. "This is the beginning of an exciting era of developmental biology and approaches like ours promise to help reveal how emerging cell types are defined at the beginning of plant life," concludes Michael Nodine with a confident smile of satisfaction.
More information: Ping Kao et al, Gene expression variation in arabidopsis embryos at single-nucleus resolution, Development (2021). DOI: 10.1242/dev.199589
Provided by Gregor Mendel Institute of Molecular Plant Biology of the Austrian Academy of Sciences