Who's in this ocean? Tracking down species on the go using environmental DNA

Sloughed off skin and bodily fluids are things most people would prefer to avoid.
But for marine biologists like Cheryl Lewis Ames, Associate Professor of Applied Marine Biology in the Graduate School of Agricultural Science at Tohoku University (Japan), such remnants of life have become a magical key to detecting the unseen.
Any organism living in the ocean will inevitably leave behind traces containing their DNA—environmental DNA (eDNA)—detectable in water samples collected from the ocean
Only recently has molecular sequencing technology become advanced enough to conduct eDNA analysis in the field to identify species that may be endangered, invasive or dangerous, and could otherwise go unnoticed.
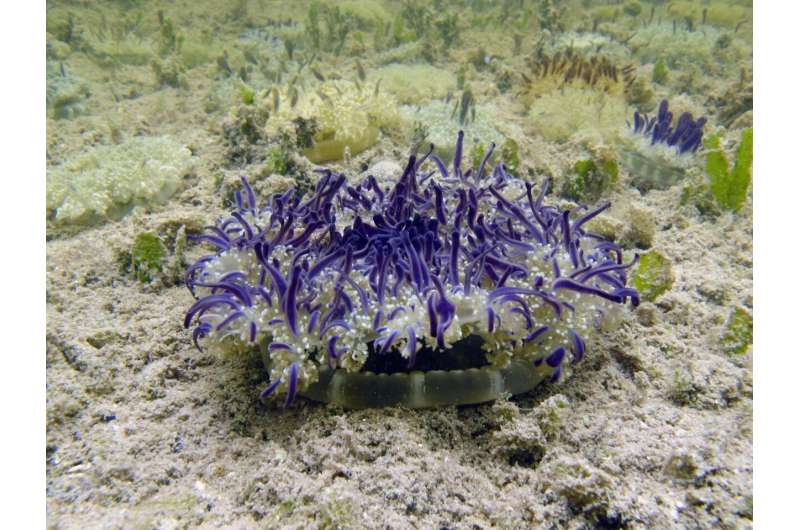
Ames chose sampling sites in the Florida Keys (U.S.) where species of upside-jellyfish Cassiopea occur to test out their newly developed Fieldable eDNA sequencing kit—called FeDS.
Using eDNA for multiple species identification is a multistep process known as metabarcoding. Since the mixed DNA template must first be amplified using the polymerase chain reaction (PCR), metabarcoding experiments in the field have only recently been possible thanks to battery-operated thermocyclers and other portable devices.

Determining the identity of a species from small bits of DNA filtered from seawater requires the use of a Next Generation Sequencer (NGS), a machine that traditionally takes up the whole desktop counter in a laboratory and requires an electrical outlet. A novelty of the portable technology used in this study—Nanopore MinION—is that as pieces of DNA pass through a microscopic pore in the device, differences in electrical current determine the unique code of each DNA strand.
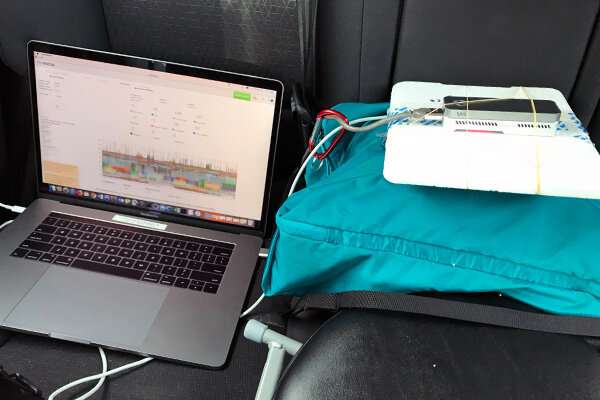
Instead of the typically large sequencing machines used to complete such a task, Ames and her team were testing a Nanopore system the size of a cellphone, powered by a laptop computer. A drop of the prepared eDNA mixture added to the portable sequencer reveals on the screen the genetic code of all the DNA passing through it in real-time.
Then, DNA sequences are searched against a huge database of sequences to determined which species are represented by the eDNA collected onsite that day. Upgraded "offline" versions of the necessary software and pre-downloading of the reference database to the laptop meant that the whole metabarcoding process could be conducted beyond the walls of a laboratory, away from internet connection.
Ames and the team detected 53 species of jellyfish including Cassiopea, the upside-down jellyfish, two venomous species of box jellyfish, many species with hydroid forms, and two species of stalked jellyfish which were previously unreported in the Florida Keys, indicating that the process could reveal species that would otherwise go unnoticed.
"My hope is that one day this system is used for sting mitigation, almost like a weather forecast app that also reports 'jellyfish stings risk' at certain beaches," said Ames.
Ames has spent much of her time conducting research in areas where jelly stings are common, and warnings about whether venomous jellies are in the area could prevent countless injuries to swimmers. Besides practical purposes in fisheries and conservation, the fact that a sample of ocean water can reveal the organisms in the vicinity is truly a marvel.
More information: Cheryl Lewis Ames et al, Fieldable Environmental DNA Sequencing to Assess Jellyfish Biodiversity in Nearshore Waters of the Florida Keys, United States, Frontiers in Marine Science (2021). DOI: 10.3389/fmars.2021.640527
Journal information: Frontiers in Marine Science
Provided by Tohoku University



















