Development of microsatellite markers for censusing of endangered rhinoceros
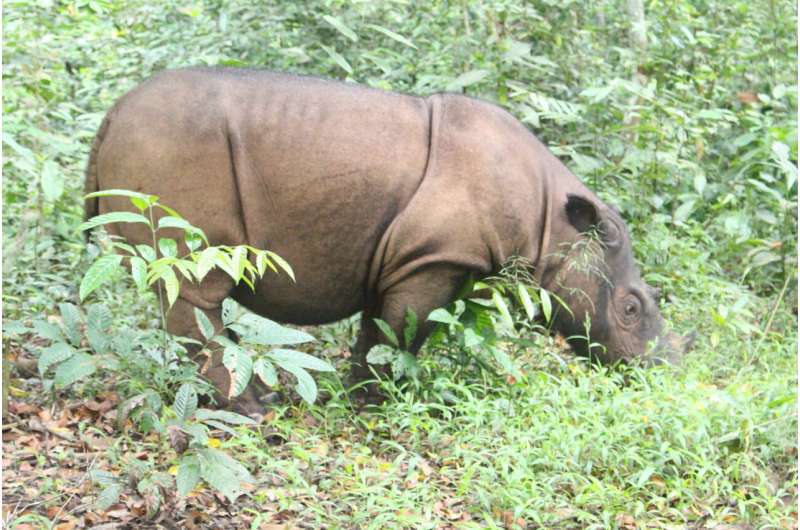

Today, the Sumatran rhinoceros (Dicerorhinus sumatrensis) is critically endangered, with fewer than 100 individuals surviving in Indonesia on the islands of Sumatra and Borneo. To ensure survival of the threatened species, accurate censusing is necessary to determine the genetic diversity of remaining populations for conservation and management plans.
A new study reported in BMC Research Notes characterized 29 novel polymorphic microsatellite markers—repetitive DNA sequences—that serve as a reliable censusing method for wild Sumatran rhinos. The study was a collaborative effort involving the University of Illinois Urbana-Champaign, the Eijkman Institute for Molecular Biology in Indonesia, Queen's University in Canada, and the San Diego Zoo.
"It's hard to do population censusing for this species because there's not a ton of them and they're very elusive so it's hard to figure out how many there are," said Jessica Brandt, former Ph.D. student in the Roca lab who led the study. "We were looking for ways to do that without handling the species. This was a joint effort between groups of people who were interested in working on these endangered species and contributing to their management."
Sumatran rhinos live in dense rainforests that are hard to traverse through, making it difficult to track Sumatran rhinoceros populations. The researchers relied on fecal DNA collected from Sumatran rhino dung samples, requiring little interaction with individuals in the wild. Although dung sampling poses many benefits, fecal DNA can be degraded and the age of the samples hard to determine. In order to overcome these challenges, researchers designed optimized microsatellite markers that were short and easy to amplify from dung samples.
"Microsatellite markers are found in non-coding regions and because of that, they evolve pretty quickly," said Brandt. "They're really useful in populations where you want to identify individuals because you're going to see more variation at those particular markers than if you're using a protein-coding gene."
"During replication of the DNA, these markers can very easily expand or contract like a genomic accordion" said University of Illinois Urbana-Champaign animal sciences professor Alfred Roca, also a member of the Carl R. Woese Institute for Genomic Biology. "By looking at enough of these markers, you can distinguish animals because microsatellites evolve very quickly and are highly variable within species. These markers are ideal to target regions of the rhino genome that are highly variable."
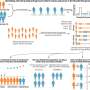
Using high quality DNA sequences from captive Sumatran rhinos, researchers identified 29 polymorphic candidate loci for further optimization. To test its utility for censusing, 13 of the 29 markers were randomly tested on fecal samples collected from wild Sumatran rhinos. The researchers were able to amplify nine of the markers from 11 wild fecal samples.
"The combination of these markers yielded better statistical power for identifying Sumatran rhino individuals and amplified very well when tested using non-invasive samples," said Sinta Saidah, co-author and research assistant at the Eijkman Institute for Molecular Biology in Indonesia. "We hope that we can use these markers on more samples collected in the field to provide island-wide population data for Sumatran rhinoceros species, which will help us devise better conservation strategies for this critically endangered species."
"To make a conservation plan, you have to know who's there and what their current level of diversity is," said Brandt. "Our markers would allow the Indonesian officials to determine not just how many rhinos they can count but whether or not they're related. Ultimately, another goal would be to expand this research to include other endangered rhinoceros species."
More information: Jessica R. Brandt et al, Characterization of 29 polymorphic microsatellite markers developed by genomic screening of Sumatran rhinoceros (Dicerorhinus sumatrensis), BMC Research Notes (2021). DOI: 10.1186/s13104-021-05522-x
Provided by University of Illinois at Urbana-Champaign