Genome of wild legume provides insights into tolerance to environmental stress
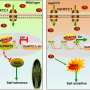
Medicago ruthenica, a wild and perennial legume forage widely distributed in semi-arid grasslands, is distinguished by its outstanding tolerance to environmental stress. It is a close relative of commonly cultivated forage of alfalfa (Medicago sativa).
The high tolerance of M. ruthenica to environmental stress makes this species a valuable genetic resource for understanding and improving traits associated with tolerance to harsh environments.
Scientists from the Institute of Botany of the Chinese Academy of Sciences (IBCAS) and their collaborators sequenced and assembled genome of M. ruthenica. They elucidated mechanisms underlying the tolerance of M. ruthenica to environmental stress by comparative genomics and transcriptomic analyses.
This study was published in the journal BMC Biology on May 6.
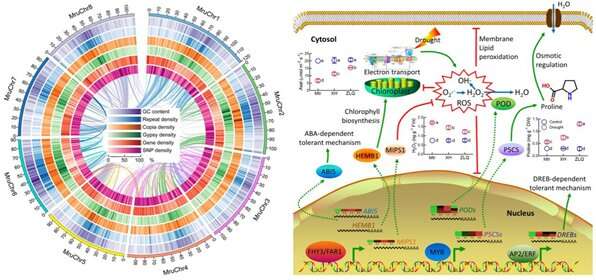
The researchers discovered that expanded FHY3/FAR1 family was involved in tolerance of M. ruthenica to drought stress. They found that many genes involved in tolerance to abiotic stress were retained in M. ruthenica compared to other cultivated Medicago species.
They identified hundreds of candidate genes associated with drought tolerance by analyzing variations in single nucleotide polymorphism using accessions of M. ruthenica with varying tolerance to drought.
Transcriptomic data revealed the involvements of genes related to transcriptional regulation, stress response and metabolic regulation in tolerance of M. ruthenica.
"The high quality genome assembly and identification of drought-related genes in the wild species of M. ruthenica provide a valuable resource for genomic studies on perennial legume forages," said Prof. ZHANG Wenhao, corresponding author of the study.
More information: Tianzuo Wang et al. The genome of a wild Medicago species provides insights into the tolerant mechanisms of legume forage to environmental stress, BMC Biology (2021). DOI: 10.1186/s12915-021-01033-0
Journal information: BMC Biology
Provided by Chinese Academy of Sciences