April 16, 2021 feature
Designed proteins assemble antibodies into modular nanocages
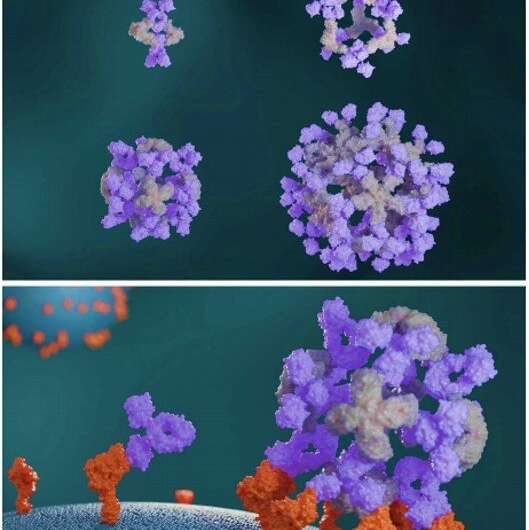
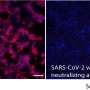
In a new report now published on Science, Robby Divine and an interdisciplinary research team at the department of biochemistry, regenerative medicine, and vaccines and infectious disease at the University of Washington U.S. and the School of Medicine, at the Tehran University of Medical Sciences, Iran, conducted computational designs of nanocages to assemble antibodies into precise architectures. During the construction, one structural component formed an antibody or Fc-ligand fusion and the second design formed an antibody-binding homo-oligomer to drive nanocage assembly with different valencies and symmetry. The team hypothesize how this process can also increase the neutralization of a pseudovirus; severe acute respiratory syndrome coronavirus-2 (SARS-COV-2) via α- SARS-COV-2 monoclonal antibodies and Fc-angiotensin-converting enzyme 2 (ACE-2) fusion proteins.
Antibodies in medical research
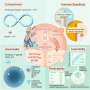
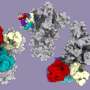
Antibodies that specifically bind to targets of interest play a central role in biomedical research and medicine. Researchers can generate clusters of antibodies by genetically linking antibody fragments together to improve signaling. It is presently difficult to form antibody assemblies with a variety of precise architectures and valencies. In this work, Divine et al. computationally designed proteins that assembled antibodies into precise architectures with different valencies and symmetries. The team hypothesized that such designs could robustly drive arbitrary antibodies into homogenous and structurally well-defined nanocages for pronounced effects on cell signaling. The researchers designed proteins to drive the assembly of arbitrary antibodies into symmetric assemblies with well-defined structures. For this, they rigidly fused together three types of "building block" units containing antibody Fc-binding domains, helical repeat connectors and cyclic oligomer-forming modules. In its architecture, the Fc-binding unit positioned itself with the C2 antibody dimer, the cyclic homo-oligomer formed the second cyclic symmetry axis in the nanocage, and the helical repeat connector linked the antibody and cyclic homo-oligomer symmetry axes in the correct orientation to form the antibody nanocages referred to as the AbCs.
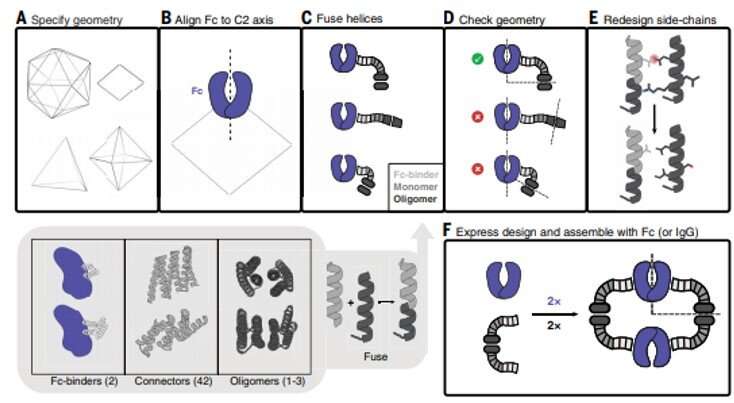
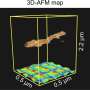
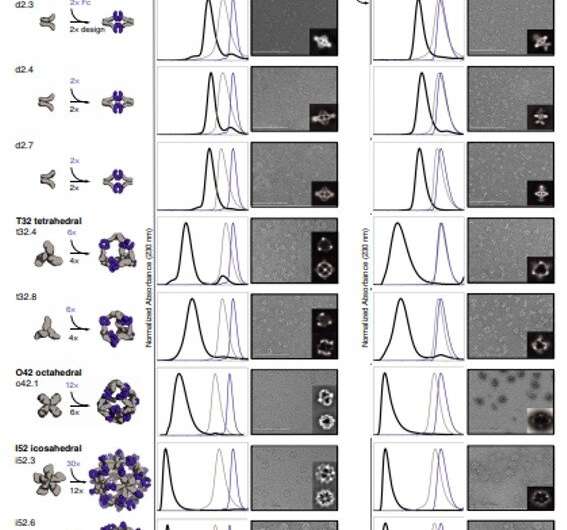
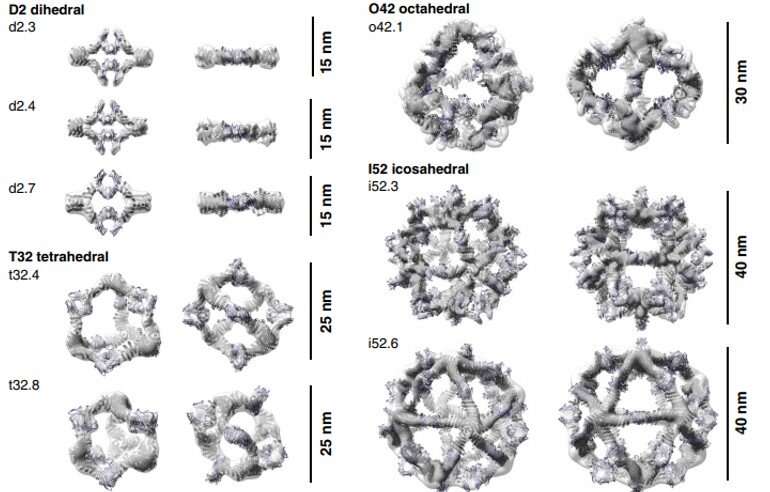
To form antibody cage (AbC)-designs, Divine et al. rigidly fused antibody constant domain-binding molecules to cyclic oligomers using helical spacer domains. They facilitated the process through helical spacer domains so that the symmetry axes of the dimeric antibody and cyclic oligomer could be at orientations that generated diverse dihedral or polyhedral architectures. The scientists optimized the junction regions between the connected building blocks to fold to the designed structures. The fusion approach relied on large sets of building blocks with many possible fusion sites per building block as stringent geometric criteria to form the desired symmetric architecture. The team used a computational method for antibody cage design to generate dihedral, tetrahedral, octahedral and icosahedral AbCs and used a naming convention to describe the final nanocage architectures. Divine et al. next expressed synthetic genes encoding the designs in bacterial cultures of Escherichia coli. For instance, the successful designs included D-2 decahedral (three designs), T-32 (one design) and 152 icosahedral (two designs) architectures containing two, six, 12 or 30 antibodies, respectively. They characterized the Fc AbCs using small-angle X-ray scattering and electron microscopy. The reconstructed nanocages were in close agreement with the computational design models. To assess the stability of nanocages, Divine et al. used dynamic light scattering readings to obtain encouraging stability to allow the characterization of their biological impact next.
Effects on cell signaling
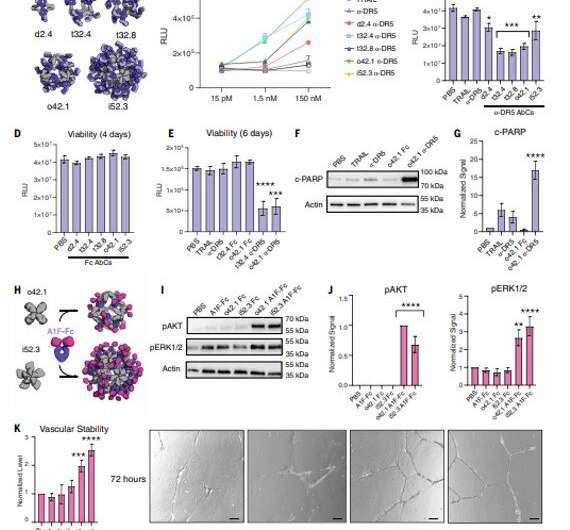
The designed AbCs provided a general platform to understand the effect of valency and geometry of receptor engagement on signaling pathway activation. The wide-range of receptor binding antibodies and natural ligands formed with the AbC method developed in this work allowed ready and systematic probing of the effect of geometry and valency of receptor subunit association on cell signaling for almost any pathway. To explore the practicality of this approach, Divine et al. assembled antibodies or Fc-ligand fusions targeting a variety of signaling pathways into Antibody cages (AbCs) and studied their impact on signaling. For instance, the AbCs, formed with a death receptor-targeting antibody induced apoptosis (cell death) of tumor cell lines that had hitherto remained unaffected by the soluble antibody or the native ligand. In addition, the assembly of Fc-fusions or antibodies in AbCs, allowed enhanced angiopoietin pathway signaling, CD-40 signaling and T-cell proliferation. The AbC formation further allowed the neutralization of an in vitro pseudovirus, such as severe acute respiratory syndrome coronavirus 2.
Outlook
The method detailed in this work, went beyond previous computational design efforts to create protein nanomaterials that integrate form and function. The AbCs therefore used antibodies as both structural and functional components to achieve a wide-range of geometries and orientations. This strategy is applicable to design vaccines with nanocages assembled with viral glycoproteins using components terminating in glycoprotein binding domains to maximize the proximity of active sites. In this way, Robby Divine and colleagues designed multiple antibody cage forming proteins to accurately cluster any protein-binding antibody into nanocages through controlled valency and geometry. The team used two, six and 12 or 30 antibodies within the AbCs by simply mixing the antibody with the corresponding designed protein without additional covalent modifications. The scientists added receptor-binding or virus-neutralizing antibodies into ABCs to enhance their biological activity across varied cell systems. The team expect this outcome for rapid antibody assembly within ordered nanocages without covalent modifications to have broad applications across research and medicine.
More information: Divine R. et al. Designed proteins assemble antibodies into modular nanocages, Science, 10.1126/science.abd9994
Ueda G. et al. Tailored design of protein nanoparticle scaffolds for multivalent presentation of viral glycoprotein antigens, eLife, 10.7554/eLife.57659
Graille M. et al. Crystal structure of a Staphylococcus aureus protein A domain complexed with the Fab fragment of a human IgM antibody: Structural basis for recognition of B-cell receptors and superantigen activity, PNAS, doi.org/10.1073/pnas.97.10.5399
Journal information: Science , Proceedings of the National Academy of Sciences , eLife
© 2021 Science X Network