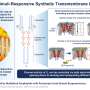
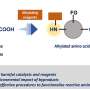
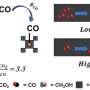
Under pressure: Manipulating protein-mimicking molecules with hydrostatic pressure
Stimulus-responsive supramolecular structures have emerged as an alternative to conventional ones, owing to their applications in sensing, drug delivery, and switchable memory systems. Now, scientists at Tokyo Institute of Technology explore the hydrostatic-pressure response of 'foldamers'—artificial molecules that mimic protein folding—and report a shift in their preferred conformation with changing pressure, demonstrating hydrostatic pressure-enabled dynamic control. The finding opens doors to future development of pressure-sensitive foldamers and artificial materials.
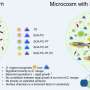
Most, if not all, biological systems are extremely complex and often rely on interactions traditional chemistry does not focus on. An entire field of research called supramolecular chemistry has been inspired to study exactly those interactions that govern biological processes, based on an approach relying on artificial molecular machines to mimic biological functions. These molecular machines are responsive to a wide range of external stimuli, such as temperature, surrounding media, excitation with light, and consequently find applications in sensing, drug delivery, molecular imaging, and switchable memory technology.
However, a particular stimulus—namely, hydrostatic pressure—has long been in fashion due to the remarkable fact that it allows a supramolecular material to be studied both in its unperturbed and pressurized state. In fact, a research group based in Japan, consisting of scientists from Tokyo Institute of Technology (Tokyo Tech), have recently shown that the optical properties and chemical processes in solutions of supramolecular materials can be precisely regulated by hydrostatic pressure.
Inspired by their findings, the group, led by Prof. Gaku Fukuhara from Tokyo Tech and Prof. Hiromitsu Maeda from Ritsumeikan University, moved on to study the effects of pressure on 'foldamers'—artificial molecules that mimic proteins. Their findings were published in the journal Chemical Science. The name foldamer derives from the fact that these systems can replicate proteins folding into well-defined patterns. Prof. Gaku Fukuhara explains the motivation for their study: "The solution-state behavior of foldamers under hydrostatic pressure has not been examined in detail, which poses a challenge to further advancements in supramolecular chemistry."
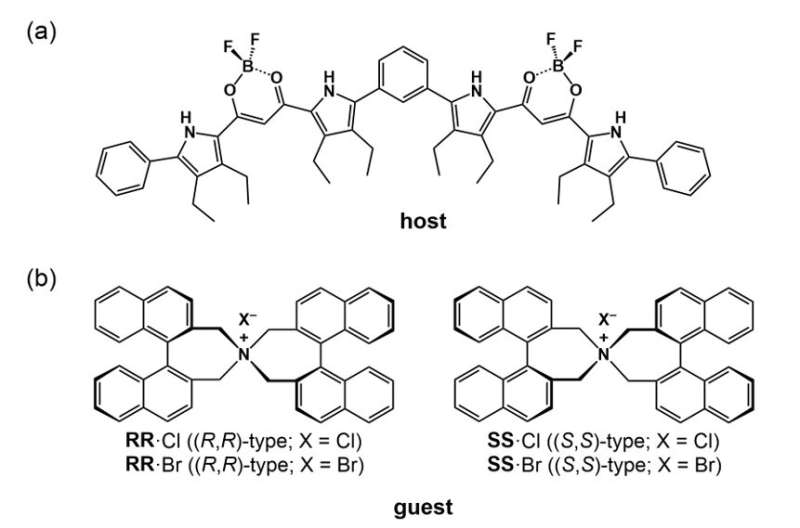
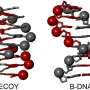
For a foldamer to fold into a specific conformation, it first needs to bind to a guest negatively charged ion that forms the racemic state (equal amount) of helical structures. The chirality (or the property of being distinct from its mirror image) of the resulting structure can then be induced by introducing an asymmetric counter positive ion, a process known as ion pairing. The ion pairing, however, depends on the solvation conditions for the foldamer, which, in turn, can be influenced by hydrostatic pressure. Accordingly, scientists chose a fluorescent foldamer as a negative ion receptor, called a host, and chiral ion pairs (such as chloride and bromide) as guests to explore the optical properties of the host solution under hydrostatic pressure.
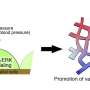
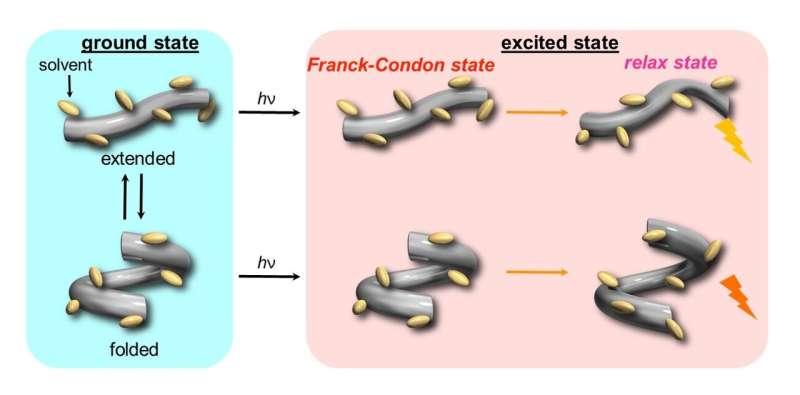
The scientists began by examining the changes in fluorescence and absorption (in UV and visible) spectra for the host in various organic solvents under pressure and observed a gradual shift in the spectral band to longer wavelengths as well as an increased absorbance with rising pressure. They attributed this to the fact that the host initially adopts two conformers, one extended and one folded, and gradually shifts to an extended-rich state upon pressurization. Then, after electronic excitation (hυ), two different fluorescence states were observed from these conformers.
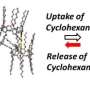
"Our study clearly shows that conformations in the flexible foldamer host can be dynamically controlled, in both ground and excited states, simply by changing hydrostatic pressure," comments an excited Prof. Fukuhara. "In fact, this strategy can even be extended to other foldamer and guest combinations that have difficulty sensing each other or do not show a host-guest chemistry," he adds.
The team's efforts at deciphering foldamers better certainly brings us one step closer to comprehending the complexity of proteins.
More information: Tomokazu Kinoshita et al, Ground- and excited-state dynamic control of an anion receptor by hydrostatic pressure, Chemical Science (2021). DOI: 10.1039/D1SC00664A
Journal information: Chemical Science
Provided by Tokyo Institute of Technology