Uncovering the structure and gating mechanism of a7 protein
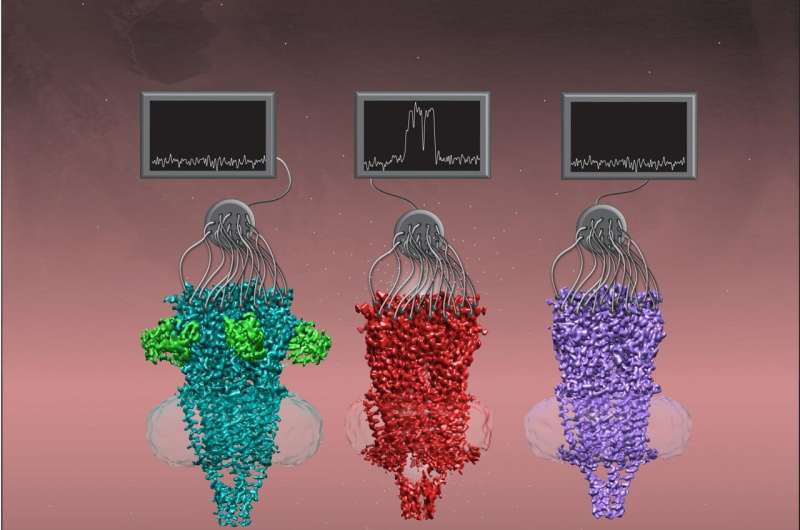

UT Southwestern researchers have identified the structure of a key member of a family of proteins called nicotinic acetylcholine receptors in three different shapes. The work, published online today in Cell, could eventually lead to new pharmaceutical treatments for a large range of diseases or infections including schizophrenia, lung cancer, and even COVID-19.
Nicotinic acetylcholine receptors are members of a broader super-family of proteins called Cys-loop receptors that function as ion channels on cell surfaces and are found in the membranes of many cell types. When the right molecule settles on these receptors, it opens the gated channels, letting ions flood from the outside to the inside of cells to trigger other cellular processes. Nicotinic acetylcholine receptors respond to acetylcholine, a molecule that nerve cells use to communicate with each other. However, they also respond to other molecules found outside the body, such as nicotine, the essential nutrient choline, and a toxin found on the skin of poison dart frogs called epibatidine.
These receptors, which have been identified in nerve, lung, and immune cells, are connected to conditions such as mental illness, neurodegenerative diseases, lung cancers, and even the destructive immune reactions characteristic of COVID-19 and other infections.
Researchers had previously identified the structures of some members of the Cys-loop super-family, a significant step toward creating drugs that fit onto these ion channels to block or enhance their function. However, say study leaders Colleen M. Noviello, Ph.D., Senior Research Scientist at UTSW, and Ryan E. Hibbs, Ph.D., Associate Professor of Neuroscience and Biophysics at UTSW, the shape, or conformation, of these channels isn't fixed. Cys-loop receptors cycle through three major states during their gating cycle that correspond to when they're closed and waiting to respond to a ligand, or activating molecule (resting state); when they've responded to the ligand and opened for ion flow (open state); or when they are still holding the ligand but have closed again (desensitized state).
After striking out for years to characterize the structure of a key nicotinic acetylcholine receptor called a7, Noviello and Hibbs got a major boost in 2016 after UTSW purchased equipment for cryogenic electron microscopy, or cryo-EM. This technology allows scientists to take photos of biologic molecules at atomic level resolution. When the researchers and their colleagues added to a7 a chaperone protein—a protein that helps support and protect other proteins—it helped a7 retain its proper shape.
With these new tools, the researchers imaged a7 in its different conformations. Because a7's structure is dynamic, allowing it to shift and wiggle, the researchers added different ligands to stabilize it so the cryo-EM images wouldn't be blurry from motion.
When they analyzed these images, Noviello, Hibbs, and their colleagues found several interesting structural features that helped explain some of a7's unusual properties. For example, a negatively charged amino acid in a7's inner channel seems to draw in positively charged calcium from outside cells, explaining why a7 is so permeable to calcium. Also, a curved structure on a7 seems to function as a latch to open the gated channel.
The team says that these newly identified structures for a7 could eventually be used as a template for pharmaceutical companies to develop new medicines that target this and related nicotinic acetylcholine receptors. They plan to continue studying a7 in various cell types and how it interacts with other molecules and proteins.
"The more we know about this important receptor found on so many diverse cell types, the closer we'll get to understanding how it functions in physiology and disease," says Hibbs, an Effie Marie Cain Scholar in Medical Research and member of the Peter O'Donnell Jr. Brain Institute.
More information: Colleen M. Noviello et al, Structure and gating mechanism of the α7 nicotinic acetylcholine receptor, Cell DOI: doi.org/10.1016/j.cell.2021.02.049
Md. Mahfuzur Rahman et al. Structure of the Native Muscle-type Nicotinic Receptor and Inhibition by Snake Venom Toxins, Neuron (2020). DOI: 10.1016/j.neuron.2020.03.012
Provided by UT Southwestern Medical Center




















