How Streptococcus pyogenes can survive on skin and cause skin infections
Streptococcus pyogenes is one of the most important bacterial causes of human skin infections. If S. pyogenes invades deep into the tissue, it can cause life-threatening illnesses, such as sepsis and toxic shock. With its limited supply of carbohydrates, the skin is generally an effective barrier against infection and not a good surface for the survival of S. pyogenes. To survive successfully and invade deep into the tissue, bacteria must be able to find a source of nutrients and also evade the skin's immune defenses.
Now, an international team led by Osaka University, Japan, in collaboration with Keio University, Japan, and University of California San Diego, U.S., has discovered a way this disease organism obtains nutrition from the skin surface. This knowledge could lead to new therapeutic approaches to tackle infections. The team recently published the work in Cell Reports.
It was already known that some bacteria break down arginine (an amino acid—one of the building blocks of proteins) via a biochemical pathway named the arginine deaminase (ADI) pathway. The team confirmed that S. pyogenes can survive using arginine even when starved of glucose. When they deleted the bacterial gene (named arcA) encoding the first enzyme in the ADI pathway, the S. pyogenes lost that ability and also became less toxic towards human skin cells grown in culture.
When arginine is supplied, the ADI pathway of S. pyogenes is activated but there are also widespread changes in gene expression. Some genes are down-regulated but large numbers of genes are up-regulated, including genes associated with virulence of S. pyogenes, such as those that produce bacterial toxins.
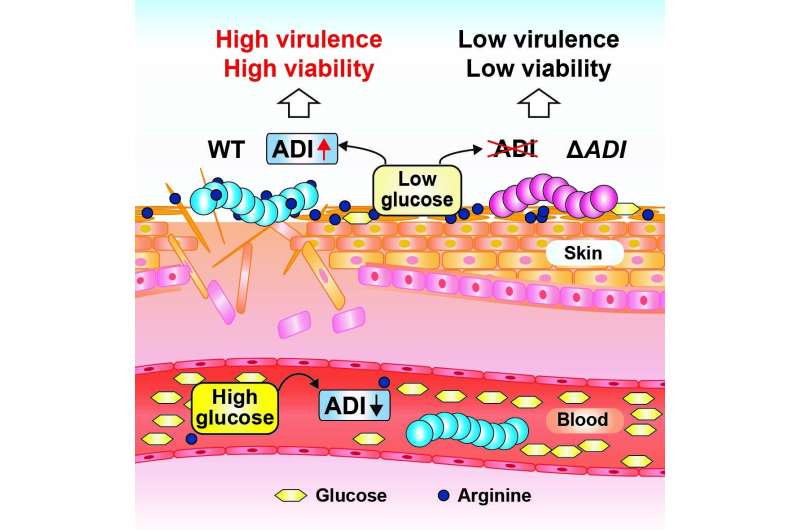
Using mouse skin as a model system for human skin, the team showed for the first time that S. pyogenes can use arginine to survive on the skin surface. S. pyogenes lacking the arcA gene survived poorly and were less virulent on mouse skin, confirming the importance of this pathway in the disease process. In contrast, in blood (containing plentiful glucose), the altered S. pyogenes appeared no different to normal S. pyogenes since the ADI pathway was not needed.
"We showed that arginine from stratum corneum-derived filaggrin was a key substrate for the ADI pathway of S. pyogenes," says lead researcher Dr. Yujiro Hirose. "In mice that do not produce filaggrin, less arginine is available and the S. pyogenes bacteria do not utilize ADI pathway to cause the skin lesion."
"This represents a significant step forward in understanding how S. pyogenes survives on the skin," explains Dr. Hirose. "We expect our study will lead to novel treatment strategies: if we can block arginine metabolism in S. pyogenes it should restrict the pathogen's survival and virulence."
More information: Yujiro Hirose et al. Streptococcus pyogenes upregulates arginine catabolism to exert its pathogenesis on the skin surface, Cell Reports (2021). DOI: 10.1016/j.celrep.2021.108924
Journal information: Cell Reports
Provided by Osaka University