Twinkle, twinkle, little star: Physicists develop a new method for counting molecules
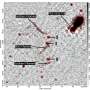
Who hasn't looked at the sky on a mild summer night and thought about the vastness of the universe? The trained eye can see the Andromeda galaxy as a distant spot. Thanks to the latest telescopes, we know that it consists of over a trillion stars. In the "nanocosm," clusters of individual light sources, such as molecules, also appear as points.
Resolving these light sources spatially is the motivation behind the field of ultrafast nanoscopy, a goal pursued in the new Regensburg Center for Ultrafast Nanoscopy RUN, a largescale interdisciplinary research facility set to open its gates in 2023.Scientists around Dr. Jan Vogelsang from the University of Regensburg, Dr. Gordon Hedley (University of Glasgow) and Professor Philip Tinnefeld (LMU Munich) now report in Nature Communications how to count the number of molecules in such a "nanostar."
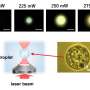
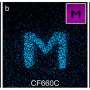
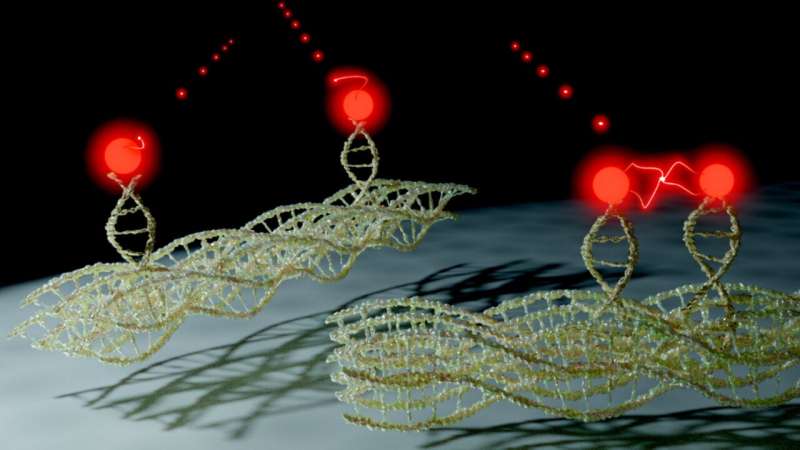
The team placed individual dye molecules at well-defined distances from one another. This is achieved by using a new method known as 'DNA origami.' DNA as the information storage medium of biology is used and programmed in such a way that the molecules are arranged by folding the DNA as desired at sizes of a few nanometres.
At first, the fluorescence light of the individual molecules on the origami cannot be distinguished under the light microscope. To actually resolve the molecules, another trick is used. The light from the origami structure is passed through a semi-transparent mirror and recorded by photodetectors on both sides of the mirror.
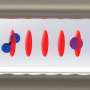
It should be noted that a single molecule can only emit a single light particle per point in time, which is only recorded by one or the other detector, but not by both. By considering the chronological order in which the light hits the individual detectors, it is possible to infer the exact number of dye molecules in the origami structure.
In this way, individual dye molecules can be counted. The number of dye molecules is determined by the programming the DNA. An origami structure with a single dye emits exactly one light quantum, one with five emits precisely five.
The individual dye molecules also interact with one another accordingly. When exposed to light, the dye absorbs energy. It can either emit this again in the form of light or pass it on to a neighboring dye. If, however, the neighboring dye is already in an excited state, two excitations will meet.
As in the case of two cars trying to drive into a single parking lot at the same time, the excitation is destroyed. Such 'annihilation' is of great importance in molecular optoelectronics such as in OLEDs or solar cells, but also plays a role in super-resolution microscopy.
The research group around Dr. Vogelsang has now been able to show that the nanoscopic interaction of the dye molecules with one another can be followed directly by resolving the arrival times of the light particles on the two light detectors. This approach offers a novel method of ultrafast nanoscopy of molecular complexes that will also find applications in the life sciences.
More information: Gordon J. Hedley et al. Picosecond time-resolved photon antibunching measures nanoscale exciton motion and the true number of chromophores, Nature Communications (2021). DOI: 10.1038/s41467-021-21474-z
Journal information: Nature Communications
Provided by University of Regensburg