A powerful computational tool for efficient analysis of cell division 4-D image data
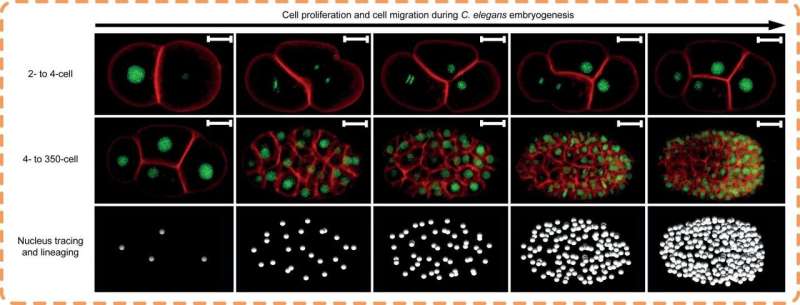
A joint research team co-led by City University of Hong Kong (CityU) has developed a novel computational tool that can reconstruct and visualize three-dimensional (3-D) shapes and temporal changes of cells, speeding up the analyzing process from hundreds of hours by hand to a few hours by the computer. Revolutionizing the way biologists analyze image data, this tool can advance further studies in developmental and cell biology, such as the growth of cancer cells.
The interdisciplinary study was co-led by Professor Yan Hong, Chair Professor of Computer Engineering and Wong Chung Hong Professor of Data Engineering in the Department of Electrical Engineering (EE) at CityU, together with biologists from Hong Kong Baptist University (HKBU) and Peking University. Their findings have been published in the scientific journal Nature Communications, titled "Establishment of a morphological atlas of the Caenorhabditis elegans embryo using deep-learning-based 4-D segmentation."
The tool developed by the team is called "CShaper." "It is a powerful computational tool that can segment and analyze cell images systematically at the single-cell level, which is much needed for the study of cell division, and cell and gene functions," described Professor Yan.
Bottleneck in analyzing massive amount of cell division data
Biologists have been investigating how animals grow from a single cell, a fertilized egg, into organs and the whole body through countless cell divisions. In particular, they want to know the gene functions, such as the specific genes involved in cell divisions for forming different organs, or what causes the abnormal cell divisions leading to tumourous growth.
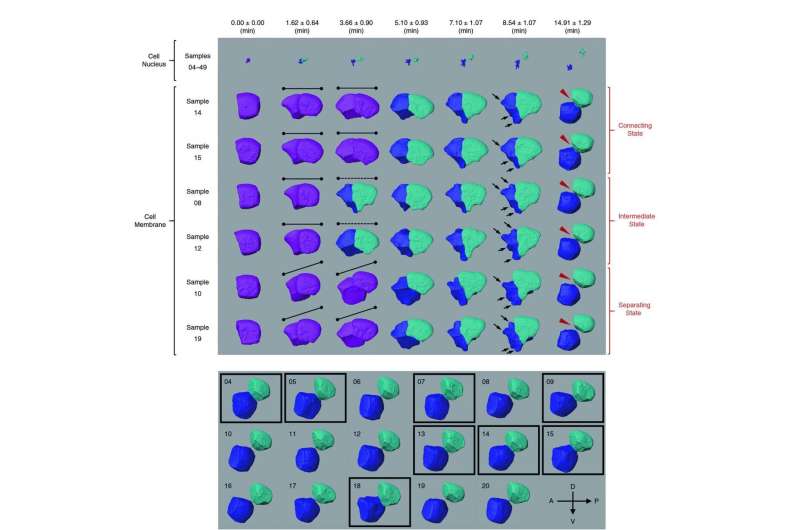
A way to find the answer is to use the gene knockout technique. With all genes present, researchers first obtain cell images and the lineage tree. Then they "knock out" (remove) a gene from the DNA sequence, and compare the two lineage trees to analyze changes in the cells and infer gene functions. Then they repeat the experiment with other genes being knocked out.
In the study, the collaborating biologist team used Caenorhabditis elegans (C. elegans) embryos to produce terabytes of data for Professor Yan's team to perform computational analysis. C. elegans is a type of worm which share many essential biological characteristics with humans and provide a valuable model for studying the tumor growth process in humans.
"With estimated 20,000 genes in C. elegans, it means nearly 20,000 experiments would be needed if knocking out one gene at a time. And there would be an enormous amount of data. So it is essential to use an automated image analysis system. And this drives us to develop a more efficient one," he said.
Breakthrough in segmenting cell images automatically
Cell images are usually obtained by laser beam scanning. The existing image analysis systems can only detect cell nucleus well with a poor cell membrane image quality, hampering reconstruction of cell shapes. Also, there is a lack of reliable algorithm for the segmentation of time-lapsed 3-D images (i.e. 4-D images) of cell division. Image segmentation is a critical process in computer vision that involves dividing a visual input into segments to simplify image analysis. But researchers have to spend hundreds of hours labeling many cell images manually.
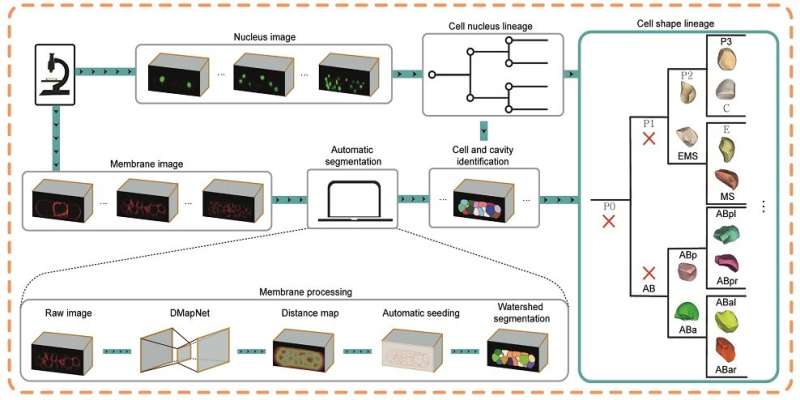
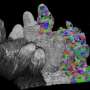
The breakthrough in CShaper is that it can detect cell membranes, build up cell shapes in 3-D, and more importantly, automatically segment the cell images at the cell level. "Using CShaper, biologists can decipher the contents of these images within a few hours. It can characterize cell shapes and surface structures, and provide 3-D views of cells at different time points," said Cao Jianfeng, a Ph.D. student in Professor Yan's group, and a co-first author of the paper.
To achieve this, the deep-learning-based model DMapNet developed by the team plays a key role in the CShaper system. "By learning to capture multiple discrete distances between image pixels, DMapNet extracts the membrane contour while considering shape information, rather than just intensity features. Therefore CShaper achieved a 95.95% accuracy of identifying the cells, which outperformed other methods substantially," he explained.
With CShaper, the team generated a time-lapse 3-D atlas of cell morphology for the C. elegans embryo from the 4- to 350-cell stages, including cell shape, volume, surface area, migration, nucleus position and cell-cell contact with confirmed cell identities.
Advancing further studies in tumor growth
"To the best of our knowledge, CShaper is the first computational system for segmenting and analyzing the images of C. elegans embryo systematically at the single-cell level," said Mr Cao. "Through close collaborations with biologists, we proudly developed a useful computer tool for automated analysis of a massive amount of cell image data. We believe it can promote further studies in developmental and cell biology, in particular in understanding the origination and growth of cancer cells," Professor Yan added.
They also tested CShaper on plant tissue cells, showing promising results. They believe the computer tool can be adopted to other biological studies.
More information: Jianfeng Cao et al, Establishment of a morphological atlas of the Caenorhabditis elegans embryo using deep-learning-based 4D segmentation, Nature Communications (2020). DOI: 10.1038/s41467-020-19863-x
Journal information: Nature Communications
Provided by City University of Hong Kong


















